Professional Documents
Culture Documents
IR Frequencies
IR Frequencies
Uploaded by
SivaPrasadCopyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
IR Frequencies
IR Frequencies
Uploaded by
SivaPrasadCopyright:
Available Formats
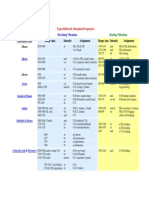
Spectroscopy Data Tables
Infrared Tables (short summary of common absorption frequencies)
The values given in the tables that follow are typical values. Specific bands may fall over a range of
wavenumbers, cm-1. Specific substituents may cause variations in absorption frequencies. Absorption
intensities may be stronger or weaker than expected, often depending on dipole moments. Additional
bands may confuse the interpretation. In very symmetrical compounds there may be fewer than the
expected number of absorption bands (it is even possible that all bands of a functional group may
disappear, i.e. a symmetrically substituted alkyne!). Infrared spectra are generally informative about what
functional groups are present, but not always. The 1H and 13C NMRs are often just as informative about
functional groups, and sometimes even more so in this regard. Information obtained from one
spectroscopic technique should be verified or expanded by consulting the other spectroscopic techniques.
IR Summary - All numerical values in the tables below are given in wavenumbers, cm-1
Bonds to Carbon (stretching wave numbers)
sp3 C-X single bonds
sp2 C-X single bonds
C
C
C
1050-1150
alkoxy C-O
1000-1350
not very useful
not used
sp2 C-X double bonds
1600-1680
not very useful
1640-1810
expanded table
on next page
1640-1690
1100-1350
acyl and phenyl C-O
1250
2100-2250
2240-2260
Stronger dipoles produce more intense IR bands and weaker dipoles produce less intense IR bands (sometimes none).
Bonds to Hydrogen (stretching wave numbers)
C
H
C
3000-3100
sp3 C-H
(see sp2 C-H bend
patterns below)
2850-3000
sp3 C-H
2700-2760
2800-2860
aldehyde C-H
(two bands)
3300
sp3 C-H
(sp C-H bend 620)
H
C
O
C
3100-3500
primary NH2
(two bands)
3100-3500
secondary N-H
(one band)
amides = strong, amines = weak
Z:\files\classes\spectroscopy\typical spectra charts.DOC
sp C-X triple bonds
C
3200-3400
2500-3400
alcohol O-H
acid O-H
2550 -2620
(very weak)
thiol S-H
Spectroscopy Data Tables
Carbonyl Highlights (stretching wave numbers)
Aldehydes
Ketones
Esters
Acids
O
C
H
saturated = 1725
conjugated = 1690
aromatic = 1700
Anhydrides
Amides
saturated = 1715
conjugated = 1690
aromatic = 1690
nitro
Acid Chlorides
O
NR2
saturated = 1650
conjugated = 1660
aromatic = 1660
6 atom ring = 1670
5 atom ring = 1700
4 atom ring = 1745
3 atom ring = 1850
saturated = 1760, 1820
conjugated = 1725, 1785
aromatic = 1725, 1785
6 atom ring = 1750, 1800
5 atom ring = 1785, 1865
alkene substitution
pattern
Cl
saturated = 1800
conjugated = 1770
aromatic = 1770
N
O
asymmetric = 1500-1600
symmetric = 1300-1390
Very often there is a very weak C=O overtone at approximately 2 x (3400 cm-1).
Sometimes this is mistaken for an OH or NH peak.,
sp2 C-H bend patterns for alkenes
descriptive
alkene term
sp2 C-H bend patterns for aromatics
absorption
frequencies (cm-1)
due to sp2 CH bend
descriptive
aromatic term
aromatic substitution
pattern
absorption
frequencies (cm-1)
due to sp2 CH bend
H
C
monosubstituted
alkene
R
C
H
C
H
C
R
C
R
C
monosubstituted
aromatic
675-730
(broad)
ortho disubstituted
aromatic
trans disubstituted
alkene
960-990
geminal disubstituted
alkene
880-900
trisubstituted
alkene
790-840
tetrasubstituted
alkene
985-1000
900-920
690-710
730-770
cis disubstituted
alkene
saturated = 1735
conjugated = 1720
aromatic = 1720
6 atom ring = 1735
5 atom ring = 1775
4 atom ring = 1840
saturated = 1715
conjugated = 1680
aromatic = 1690
6 atom ring = 1715
5 atom ring = 1745
4 atom ring = 1780
3 atom ring = 1850
R'
none
Z:\files\classes\spectroscopy\typical spectra charts.DOC
735-770
meta disubstituted
aromatic
para disubstituted
aromatic
680-725
750-810
880-900 (sometimes)
790-840
Aromatic compounds have characteristic weak overtone bands
that show up between 1650-2000 cm-1). Some books provide
pictures for comparison (not here). A strong C=O peak will
cover up most of this region.
Spectroscopy Data Tables
3
units = cm-1
4000
2500
3000
3500
sp C-H
stretch
1700
2000
sp3 C-H
stretch
C C
thiol S-H
stretch
sp2 C-H
stretch
1500 1400 1300 1200 1100 1000 900 800 700
C=O
stretch
aldehyde C-H
stretch
geminal
acyl C-O
phenol C-O
tri
aromatic sp2 C-H
bend
mono
N-H bend
ortho
2o N-H
stretch
nitro
meta
nitro
para
3000
2500
1700
2000
1500 1400 1300 1200 1100 1000 900
expansion of alkene & aromatic sp2 C-H bend region (units = cm-1)
700
600
800
900
mono
cis
trans
C=C stretch
aromatic
1o N-H2
stretch
1000
mono
alkoxy C-O
carboxylic acid O-H
stretch
3500
sp C-H
bend
C=C stretch
alkene
alcohol O-H
stretch
4000
alkene sp2 C-H
bend
C=N
stretch
C N
600 500
800 700
600 500
500
mono
cis
alkene sp2 C-H
bend
trans
geminal
tri
mono
mono
aromatic sp2 C-H
bend
ortho
meta
meta
meta
para
1750
1800
Saturated C=O lies at
higher cm-1
C=O in samll rings
lies at higher cm-1
expansion of carbonyl (C=O) stretch region (units = cm-1)
1700
1650
carboxylic acid C=O (also acid "OH")
ester C=O (also acyl C-O and alkoxy C-O)
aldehyde C=O (also aldehyde C-H)
ketone C=O (nothing special)
acid chloride C=O (high C=O, 1 peak)
anhydride C=O
anhydride C=O (high C=O, 2 peaks)
Z:\files\classes\spectroscopy\typical spectra charts.DOC
amide C=O (low C=O, amide N-H)
1600
Conjugated C=O
lies at lower cm-1
Spectroscopy Data Tables
IR Flowchart to determine functional groups in a compound (all values in cm-1).
IR Spectrum
has C=O band
(1650-1800 cm-1)
very strong
does not have
C=O band
C
aldehydes
O
1725-1740 (saturated)
1660-1700 (unsaturated)
C
sometimes lost
2860-2800
in sp3 CH peaks
2760-2700
aldehyde C-H
(both weak)
ketones
1710-1720 (saturated)
1680-1700 (unsaturated)
1715-1810 (rings: higher
in small rings)
esters - rule of 3
O
1735-1750 (saturated)
1715-1740 (unsaturated)
1735-1820 (higher in small rings)
acyl
O
1150-1350 (acyl, strong)
alkoxy
(1000-1150, alkoxy, medium)
acids
O
1700-1730 (saturated)
1715-1740 (unsaturated)
1680-1700 (higher in small rings)
acyl
C
1210-1320 (acyl, strong)
2400-3400, very broad
(overlaps C-H stretch)
acid
O
nitriles
C
a little lower
when conjugated
2150
(variable intensity)
not present or weak when symmetrically
substituted, a little lower when conjugated
C
sp C-H stretch
sp C-H bend
3300
sharp, strong
620
All IR values are approximate and have a range
of possibilities depending on the molecular
environment in which the functional group resides.
Resonance often modifies a peak's position
because of electron delocalization (C=O lower,
acyl C-O higher, etc.). IR peaks are not 100%
reliable. Peaks tend to be stronger (more intense)
when there is a large dipole associated with a
vibration in the functional group and weaker in
less polar bonds (to the point of disappearing in
some completely symmetrical bonds).
1460 & 1380
sp3 C-H bend
C
not useful
alkenes
sp2 C-H stretch
3000-3100
650-1000
(see table for
spectral patterns)
sp2 C-H bend
C
1600-1660
weak or not present
aromatics
sp2 C-H stretch
3050-3150
690-900 (see table),
overtone patterns
between 1660-2000
sp2 C-H bend
C
1600 & 1480
can be weak
alcohols
alcohol
O
3600-3500
1000-1260
(3o > 2o > 1o)
2550 (weak)
(easy to overlook)
alkoxy
C
Alkene sp C-H bending patterns
amines
1630-1680 (saturated)
1745 (in 4 atom ring)
C
H
N
o
N
H
2o
3350 & 3180, two bands
for 1o amides,
one band for 2o amides,
H stronger than in amines, extra
overtone sometimes at 3100
N-H bend, 1550-1640,
stronger in amides than amines
acid chlorides
1800 (saturated)
1770 (unsaturated)
Inductive pull of Cl increases the
electron density between C and O.
anhydrides
O
1760 & 1820 (saturated)
1725-1785 (unsaturated)
two strong bands
acyl
C
2850-3000
thiols
thiol
amides
alkanes
sp3 C-H stretch
alkynes
2250
sharp, stronger
than alkynes,
monosubstituted alkene (985-1000, 900-920)
geminal disubstituted (960-990)
cis disubstituted (675-730)
trans disubstituted (880-900)
trisubstituted (790-840)
tetrasubstituted (none, no sp2 C-H)
Aromatic sp2 C-H bending patterns
monosubstituted (730-770, 690-710)
ortho disubstituted (735-770)
meta disubstituted (880-900,sometimes,
750-810, 680-725)
para disubstituted (790-840)
H
N
N
1o
2o
N-H bend, 1550-1640,
stronger in amides than amines
1000-1350
(uncertain)
ethers
alkoxy
C
1120 (alphatic)
1040 & 1250 (aromatic)
nitro compounds
O
1500-1600, asymmetric (strong)
1300-1390, symmetric (medium)
There are also weak overtone bands between
1660 and 2000, but are not shown here. You
can consult pictures of typical patterns in other
reference books. If there is a strong C=O band,
they may be partially covered up.
3300 - 3500, two bands
for 1o amines, one band
o
H for 2 amines, weaker
than in amides,
carbon-halogen bonds
1150-1350 (acyl, strong)
X = F, Cl, Br, I
Z:\files\classes\spectroscopy\typical spectra charts.DOC
usually not
very useful
Spectroscopy Data Tables
5
Typical 1H and 13C NMR chemical shift values.
deshielding side = less electron rich
(inductive & resonance)
shielding side = more electron rich
(inductive & resonance)
typical proton chemical shifts
amine N-H
Carbon and/or heteroatoms without hydrogen do
not appear here, but influence on any nearby protons
may be seen in the chemical shifts of the protons.
alcohol
1
amide N-H
S C H
thiols, sulfides
2.5
N
3.0
X C H
X = F,Cl,Br,I
10
8+
6
7
PPM
alcohols
ethers
esters
5+
5
ketones
no H
15
N C
amines, amides
with & without H
180
50
O
O
C
180
C C
with & without H
N C
no H
90
70
40
S
thiols, sulfides
with & without H
160
O
C
60
-
110
125
30
epoxides
with & without H
carboxylic acids
anhydrides
esters
amides
acid chlorides
no H
alcohols,
ethers, esters
40
20
with & without H
H
aldehydes
with H
200
F 80-95
Cl 45-70
Br 35-65
I 15-45
220+
220
95
210
0.5
with & without H
halogen
simple sp3 C-H
CH > CH2 > CH3
3.3 3
typical carbon-13 chemical shifts
240
1.5 1.3
2.5
3.5
H
O C
aromatic C-H
10
thiol
SH
epoxide C-H
10
11
1.5
3+
7+
aldehyde C-H
12
2.5
benzylic C-H
carbonyl alpha C-H
alkene C-H
12
2.3
allylic C-H
carboxylic acid O-H
2.0
C H
amines
80
50
with & without H
180
180
160+
160
100-
140 PPM 120
Z:\files\classes\spectroscopy\typical spectra charts.DOC
100
simple sp3 carbon
C > CH > CH2 > CH3
with & without H
60+
80
60
40
20
Spectroscopy Data Tables
6
Calculation of chemical shifts for protons at sp3 carbons
H
C C C
Estimation of sp3 C-H chemical shifts with multiple substituent parameters for protons within 3 C's of consideration.
= directly attached substituent, use these values when the hydrogen and substituent are attached to the same carbon
= once removed substituent, use these values when the hydrogen and substituent are on adjacent (vicinal) carbons
= twice removed substituent, use these values when the hydrogen and substituent have a 1,3 substitution pattern
0.0
0.8
0.9
1.4
3.2
2.2
2.1
2.0
2.3
2.1
1.5
2.5
2.8
2.8
3.1
2.8
1.5
2.1
3.2
1.3
1.3
1.1
1.2
1.7
1.1
1.1
1.0
1.8
1.1
1.6
1.8
X = substituent
R- (alkyl)
R2C=CR- (alkenyl)
RCC- (alkynyl)
Ar- (aromatic)
F- (fluoro)
Cl- (chloro)
Br- (bromo)
I- (iodo)
HO- (alcohol)
RO- (ether)
epoxide
R2C=CRO- (alkenyl ether)
ArO- (aromatic ether)
RCO2- (ester, oxygen side)
ArCO2- (aromatic ester, oxygen side)
ArSO3- (aromatic sulfonate, oxygen)
H2N- (amine nitrogen)
RCONH- (amide nitrogen)
O2N- (nitro)
HS- (thiol, sulfur)
RS- (sulfide, sulfur)
OHC- (aldehyde)
RCO- (ketone)
ArCO- (aromatic ketone)
HO2C- (carboxylic acid)
RO2C- (ester, carbon side)
H2NOC- (amide, carbon side)
ClOC- (acid chloride)
NC- (nitrile)
RSO- (sulfoxide)
RSO2- (sulfone)
0.0
0.2
0.3
0.4
0.5
0.5
0.7
0.9
0.3
0.3
0.4
0.4
0.5
0.5
0.5
0.4
0.2
0.3
0.8
0.4
0.4
0.4
0.3
0.3
0.3
0.3
0.3
0.4
0.4
0.5
0.5
0.0
0.1
0.1
0.1
0.2
0.2
0.2
0.1
0.1
0.1
0.1
0.2
0.3
0.1
0.2
0.0
0.1
0.1
0.1
0.1
0.1
0.1
0.0
0.1
0.1
0.1
0.1
0.1
0.2
0.3
0.3
Starting value and equations for CH3's
CH3 = 0.9 +
H3C
CH3 = 0.9 + ( + )
H3C C C
is the summation symbol for all substituents considered
Starting value and equation for CH2's
In a similar manner we can calculate chemical shifts
for methylenes (CH2) using the following formula
CH2 = 1.2 + ( + + )
H
H C C C
is the summation symbol for all substituents considered
Starting value and equation for CH's
In a similar manner we can calculate chemical shifts
for methines (CH) using the following formula
CH = 1.5 + ( + + )
H
C C C
is the summation symbol for all substituents considered
a. methine b. methylene
d. methyl
H3C
CH2
CH3
HO
N
CH
O
H2C
H2C
e. methylene
f. methylene
c. methyl
Calculations are generally close to
actual chemical shifts for a single
substituent, but are less reliable as
the number of substituent factors
goes up. Multiple substituent factors
tend to overestimate an actual chemical
shift.
a. methine = 1.5 + (1.4) + (2.3) + (0.2) = 5.4 ppm
actual = 5.2
d. methyl = 0.9 + (0.1) = 1.0 ppm
actual = 1.0
b. methylene = 1.2 + (1.5) + (0.4) + (0.3) = 3.4 ppm
actual = 3.0 and 3.2
e. methylene = 1.2 + (0.3) = 1.5 ppm
actual = 1.7
c. methyl = 0.9 + (1.5) = 2.4 ppm
actual = 2.6
f. methylene = 1.2. + (1.7) = 2.9 ppm
actual = 2.9
Z:\files\classes\spectroscopy\typical spectra charts.DOC
Spectroscopy Data Tables
Estimated chemical shifts for protons at alkene sp2 carbons
Substituent
HHydrogen
RAlkyl
C6H5CH 2Benzyl
X-CH2Halomethyl
(H)/ROCH2alkoxymethyl
(H)2/R2NCH2aminomethyl
RCOCH2-keto
NCCH2-cyano
R2C=CRAlkenyl
C6H5Phenyl
FFluoro
ClChloro
BrBromo
IIodo
ROakoxy (ether)
RCO2O-ester
(H)2/R2NN-amino
RCONHN-amide
O2NNitro
RSThiol
OHCAldehyde
ROCKetone
HO2CC-acid
RO2CC-ester
H2NOCC-amide
NCNitrile
geminal
cis
trans
0.0
0.0
0.0
0.5
-0.2
-0.3
0.7
-0.2
-0.2
0.7
0.1
0.0
0.6
0.0
0.0
0.6
-0.1
-0.1
Substitution relative to calculated "H"
cis
H
C C
trans
gem
(ppm) = 5.2 + gem + cis + trans
Example Calculation
gem
H
C
0.7
-0.1
-0.1
0.7
-0.1
-0.1
1.2
0.0
0.0
1.4
0.4
-0.1
1.5
-0.4
-1.0
trans = 5.2 - 0.1 = 5.1
actual = 5.1
1.1
0.2
0.1
1.1
0.4
0.6
cis = 5.2 + 0.4 = 5.7
actual = 5.6
1.1
0.8
0.9
1.2
-1.1
-1.2
2.1
-0.4
-0.6
0.8
-1.3
-1.2
2.1
-0.6
-0.7
1.9
1.3
0.6
1.1
-0.3
-0.1
1.0
1.0
1.2
1.1
0.9
0.7
0.8
1.0.
03
0.8
1.0
0.5
0.4
1.0
0.5
0.3
0.8
Z:\files\classes\spectroscopy\typical spectra charts.DOC
0.6
trans
H
cis
CH3O
gem = 5.2 + 1.4 = 6.6
actual = 6.6
bH
C
aH
c
d
H H
C
C
O C
O
e
H
C
Hf
a = 5.2 + (-0.4) = 4.8
actual = 4.9 (J = 14, 1.6 Hz)
b = 5.2 + (-0.6) = 4.6
actual = 4.6 (J = 6, 1.6 Hz)
c = 5.2 + 2.1 = 7.3
actual = 7.4 (J = 14, 6 Hz)
d = 5.2 + 0.8 = 6.0
actual = 6.2 (J = 18, 11 Hz)
e = 5.2 + 0.5 = 5.7
actual = 5.8 (J = 11, 1.4 Hz)
f = 5.2 + 1.0 = 6.2
actual = 6.4 (J = 18, 1.4 Hz)
Spectroscopy Data Tables
Estimated chemical shifts for protons at aromatic sp2 carbons
Substituent
HHydrogen
CH3Methyl
ClCH2Cholromethyl
Cl3CHalomethyl
HOCH 2Hydroxymethyl
R2C=CRAlkenyl
C6H5Phenyl
FFluoro
ClChloro
BrBromo
IIodo
HOHydroxy
ROAlkoxy
RCO 2O-ester
(H)2/R2NN-amino
RCONHN-amide
O 2NNitro
RSthiol/sulfide
OHCAldehyde
ROCKetone
HO2CC-acid
RO2CC-ester
H 2NOCC-amide
NCNitrile
ortho
meta
para
0.0
0.0
0.0
-0.2
-0.1
-0.2
0.0
0.0
0.0
0.6
0.1
0.1
-0.1
-0.1
-0.1
0.1
0.0
-0.1
1.4
0.4
-0.1
-0.3
0.0
-0.2
0.0
0.0
-0.1
0.2
-0.1
0.0
0.4
-0.2
0.9
-0.6
-0.1
-0.5
-0.5
-0.1
-0.4
-0.3
0.0
-0.1
-0.8
-0.2
-0.7
0.1
-0.1
-0.3
1.0
0.3
0.4
-0.1
-0.1
-0.2
0.6
0.2
0.3
0.6
0.1
0.2
0.9
0.2
0.3
0.7
0.1
0.2
0.6
0.1
0.2
0.4
0.2
0.3
Z:\files\classes\spectroscopy\typical spectra charts.DOC
Substitution relative to calculated "H"
meta
ortho
para
H
meta
ortho
(ppm) = 7.3 + ortho + meta + para
Example Calculation
2
H
1
CH3O
2H
H
3
CH2
4
5
H 6
H
7
1. (CH3) = 0.9 + 2.8 = 3.7
actual = 3.8
2. (2) = 7.3 + (-0.5) ortho + (-0.1) para = 6.7
actual = 6.8
3. (3) = 7.3 + (-0.2) ortho + (-0.4) para = 6.7
actual = 7.1
4. (CH 2) = 1.2 + (0.8) + (1.4) = 3.4
actual = 3.3
5. (5) = 5.2 + (0.7) gem = 5.9
actual = 5.9
6. (6) = 5.2 + (-0.2) trans = 5.0
actual = 5.1
7. (7) = 5.2 + (-0.2) cis = 5.0
actual = 5.1
Spectroscopy Data Tables
Real Examples of Combination Effects on Chemical Shifts
bond anisotropy
bond example too
0.8 shielded
(CH 2)
0.8, shielded
H
shielding cone
from bond
CH2
2.6
H
7.2
electronegativity and bond
O
O
C
10-12
H O
O
C
9.5
H
1.5
deshielded
H 3C
hydrogen
bonding
O H
H
C
O
C
15, hydrogen
bonded enol
CH3
electronegative substituent and distance from protons
O CH2CH 2CH2CH2CH 3
CH3 Cl
3.0
3.6 1.5 1.3 1.3 0.9
multiple substituents
CH4
0.2
CH3CH 2 Cl
CH3CH2CH 2 Cl
1.3
1.0
CH 3Cl
CH2Cl2
3.0
= 2.8
H 3C C
CH3CH 2Cl (CH 3)2CHCl
3.0
3.5
4.1
2.4
C H 1.9
C C H 3.0
R C
Ar
RO
ArO
RS
ArO
H
H
H
H
CCl4
6.4
C
H
5.8
Ph
3.7
H 2C
H 3C
1.3
? (oops)
H 3CH2C C
Ph (H3C)2CH C
H
C
4.2
C
H
4.0
8.2
H
7.5
H
Ph
3.5
H
6.5
O
H 2N
=?
3.0
H
H
H
H
H
H
bond anisotropy
produces deshielding Extra electron density via resonance produces
shielding effect on aromatic protons, especially
effect on aromatic
at ortho/para positions.
protons.
sp C-H
H C C H
CHCl3
2.6
alkene substituent resonance and inductive effects
0.9 1.4 2.0
O
3.8
H
H CH CH CH
H 5.0 CH O C
3
2
2
3
C C
C C
C
H
H
H
H
H
5.8
4.9
5.3
6.1
aromatic resonance and inductive effects
6.6
H
H 7.3
7.1
H
H
6.7
H
H
H 2N
H
0.9
substituents at methyl (CH3), methylene (CH2) and methine (CH)
CH3Cl
0.9
7.2
= 1.9
5.3
= 2.3
CH3CH 2CH 2CH2 Cl CH3CH 2 R
7.7
H
C O
H 3C
2.1
H
C
H
7.3
4.9
C
H
4.6
O
N
O
H
H
H
H
Withdrawal of electron density via resonance
produces deshielding effect on aromatic protons,
especially at ortho/para positions.
O
R2N H amine H = 1-5 enol H = 10-17 H O
alcohol H = 1-5
C C
phenol H = 4-10
O
O
amide H = 5-8
thiol H = 1-2.5
R C
R C
aromatic thiol H = 3-4
NH2
O H acid H = 10-13
Z:\files\classes\spectroscopy\typical spectra charts.DOC
Spectroscopy Data Tables
10
1. One nearest neighbor proton
H1
observed
proton
Ha
increasing
one neighbor
proton = Ha
increasing E (, Bo)
the ratio of these
two populations
is about 50/50 (or 1:1)
H1
perturbation(s) by
neighbor proton(s)
Eto flip proton
E2 (observed)
E1 (observed)
Bo
J1a
1
Protons in this environment have a small cancellation
of the external magnetic field, Bo, and produce a
smaller energy transition by that tiny amount.
H1
C
H1
C
J = coupling constant
small difference in
energy due to differing
neighbor's spin (in Hz)
Protons in this environment have a small
additional increment added to the external
magnetic field, Bo, and produce a higher
energy transition by that tiny amount.
N + 1 rule (N = # neighbors)
J (Hz)
# peaks = N + 1 = 1 + 1 = 2 peaks
(ppm)
2. Two nearest neighbor protons (both on same carbon or one each on separate carbons)
observed
proton
H1
two neighbor
Ha protons
the ratio of these
four populations
is about 1:2:1
Hb
H1
E1
Eto flip proton
E2
J1a
E3
Bo
H1
C
two neighbor protons are like
two small magnets that can be
arranged four possible ways
(similar to flipping a coin twice)
J1b
two equal energy
populations here
J1b
N + 1 rule (N = # neighbors)
J (Hz)
J (Hz)
# peaks = N + 1 = 2 + 1 = 3 peaks
(ppm)
3. Three nearest neighbor protons (on same carbon, or two on one and one on another, or one each on separate carbons)
H1
observed
proton
three neighbor
Ha protons
C C
the ratio of these
eight populations
is about 1:3:3:1
Hb
H1
Hc
Eto flip proton
E1
E2
Bo
H1
C C
E3
J1a
E4
J1b
three neighbor protons are like
three small magnets that can be
arranged eight possible ways
(similar to flipping a coin thrice)
three equal energy
populations at each
of middle transitions
J1c
J1b
J1c
J1c
N + 1 rule (N = # neighbors)
J (Hz)
J (Hz)
(ppm)
Z:\files\classes\spectroscopy\typical spectra charts.DOC
J (Hz)
# peaks = N + 1 = 3 + 1 = 4 peaks
Spectroscopy Data Tables
11
Splitting patterns when the N+1 rule works (common, but not always true)
= group without any coupled proton(s)
N=1
N=0
N=2
H
H
C
N=3
H
H
C
H2
C
CH
CH2
CH3
CH
CH
CH
= calc or exp
CH
d, J=7
I=1H
N=1
t, J=7
I=1H
N=2
= calc or exp
= calc or exp
s, J=none
I=1H
N=0
CH
q, J=7
I=1H
N=3
= calc or exp
N=4
N=5
H
C
H
CH
CH3
CH2
CH
CH2
CH3
CH3
CH
CH
CH
CH2
CH2
CH2
CH2
CH
sex, J=7
I=1H
N=5
qnt, J=7
I=1H
N=4
= calc or exp
= calc or exp
N=6
N=7
H
H
C
CH
CH3
CH3
H2C
CH2
CH
CH2
H2 C
CH3
H
CH3
CH3
sep, J=7
I=1H
N=6
oct, J=7
I=1H
N=7
CH3
CH2
= calc or exp
= calc or exp
N=8
H2 C
Pascal's triangle = coefficients of variable terms in binomial expansion (x + y)n, n = integer
Multiplets when the N + 1 rule works (all J values are equal).
H2C
CH3
CH3
non, J=7
I=1H
N=8
1 peak = 100%
s = singlet
d = doublet
t = triplet
1
1
1
q = quartet
qnt = quintet
sex = sextet
= calc or exp
sep = septet
o = octet
1
1
1
1
6
7
2
3
4
5
1 peak = 50%
1 peak = 25%
1
1
3
6
1 peak = 12%
1
10 10 5
1 peak = 6%
1
1 peak = 3%
1
15 20 15 6
1
21 35 35 21 7 1
relative sizes of
peaks in multiplets
(% edge peak shown)
1 peak = 1.5%
1 peak = 0.8%
Combinations or these are possible.
dd = doublet of doublets; ddd = doublet of doublet of doublets; dddd = doublet of doublet of doublet of doublets; dt = doublet of triplets
td = triplet of doublets; etc.
Z:\files\classes\spectroscopy\typical spectra charts.DOC
Spectroscopy Data Tables
12
Typical Coupling Constants
Ha
Range
Typical
0-30 Hz
14 Hz
Hb
geminal protons - can have different chemical shifts
and split one another if they are diastereotopic
Range
Typical
6-8 Hz
= dihedral
angle
Range
0-3 Hz
1 Hz
Range
Typical
0-3 Hz
1 Hz
Hb
C C
Hb
trans / allylic coupling,
notice through 4 bonds
Typical
C
7 Hz
Ha
Hb
cis / allylic coupling,
notice through 4 bonds
vicinal protons are on adjacent atoms, when freely
rotating coupling averages out to about 7 Hz
Ha Hb
C
C
Typical
Ha
Ha Hb
C
Ha
Range
0-12 Hz
7 Hz
depends on dihedral
angle, see plot of
Karplus equation
Range
Typical
0-1 Hz
0 Hz
Range
Typical
9-13 Hz
10 Hz
Range
Typical
1-3 Hz
2 Hz
Range
Typical
5-8 Hz
6 Hz
Range
Typical
2-3 Hz
2 Hz
Range
Typical
2-3 Hz
3 Hz
C
C
Hb
Ha
sp2 vicinal coupling
(different bonds)
Ha
C
Hb
C
O
sp3 vicinal aldehyde coupling
protons rarely couple through 4 chemical bonds
unless in a special, rigid shapes (i.e. W coupling)
Ha
C
Range
Typical
0-3 Hz
2 Hz
Hb
Hb
C
Range
Typical
5-11 Hz
10 Hz
C
Hb
Range
Typical
11-19 Hz
17 Hz
sp2 trans coupling (always
larger than the cis isomer)
C
C
Hb
C C
Hb
Hb
Ha
C
C C
bis-propargylic coupling
notice through 5 bonds
Range
Ha
C
Ha
sp / propargylic coupling
notice through 4 bonds
sp2 cis (acylic) coupling (always
smaller than the trans isomer)
sp vicinal aldehyde coupling
Ha
sp2 geminal coupling
Ha
Hb
Ha
4-10 Hz
Typical ortho, meta and
para coupling to
7 Hz this proton
Range
H ortho
H meta
sp2 / sp3 vicinal coupling
ortho 6-10 Hz
meta 2-3 Hz
para 0-1 Hz
Hpara
When J values are less than 1 Hz, it is often difficult to resolve them and a peak may merely appear wider and shorter.
Z:\files\classes\spectroscopy\typical spectra charts.DOC
Typical
9 Hz
2 Hz
0 Hz
Spectroscopy Data Tables
13
Similar chemical shift information presented in a different format. Remember, proton decoupled
carbons appear as singlets. When carbons are coupled to their hydrogens, carbons follow the N+1 rule.
Methyls = q, methylenes = t, methines = d, and carbons without hydrogen appear as singlets = s.
DEPT provides the same information. Carbon chemical shifts are spread out over a larger
range than proton chemical shifts (they are more dispersed), so it is less likely that two different carbon
shifts will fall on top of one another. The relative positions of various types of proton and
carbon shifts have many parallel trends (shielded protons tend to be on shielded carbons, etc.)
CH2
CH3
Simple alkane
carbons
0 - 30 ppm
(q)
20 - 40 ppm
(t)
50 - 60 ppm
CH2 N
d
10 - 50 ppm
(t)
CH2 X
sp3 carbon next to
bromine or chlorine
(X = Cl, Br)
sp carbon (alkynes)
sp2 carbon (alkenes
and aromatics)
25 - 50 ppm
(t)
60 - 80 ppm d
(d)
50 - 70 ppm d
(d)
60 - 80 ppm
(d)
50 - 70 ppm
(s)
C
70 - 90 ppm
70 - 90 ppm
(s)
C N
sp carbon (nitriles)
60 - 80 ppm
(s)
C N
110 - 125 ppm
C
180 - 210 ppm
aldehyde carbons, lower
values when conjugated
(d)
Z:\files\classes\spectroscopy\typical spectra charts.DOC
O
C
140 - 160+ ppm
sp carbon attached to an electronegative atom
(X = oxygen, nitrogen, halogen) or C carbon
conjugated with a carbonyl group
100 - 140 ppm
simple sp2 carbon
resonance donation moves lower,
resonance withdrawal moves higher
160 - 180 ppm
carboxyl carbons
(acids, esters, amides)
(s)
C O
CH X
30 - 60 ppm
(s)
CH N
35 - 55 ppm
(q)
(t)
CH3 N
30 - 50 ppm
(d)
CH O
55 - 80 ppm
(q)
sp3 carbon
next to nitrogen
CH2 O
CH3 O
sp3 carbon
next to oxygen
CH
R
180 - 220 ppm
ketone carbons, lower
values when conjugated
(s)
Spectroscopy Data Tables
14
Calculations of alkane 13C chemical shifts not listed above.
sp3 Carbon Chemical Shift Calculations
Calculations for sp3 carbon 13C chemical shifts of functionalized carbon skeletons can be performed starting
from the actual shifts found in the corresponding alkane skeleton, and introducing corrections factors based on the
functionality present in the molecule. This assumes that the alkane 13C shifts are available, which is why several
examples are provided below.
Examples of Cn alkanes as possible starting points for calculation 13C shifts in ppm.
Approximate 13C shift calculation from scratch.
Steric Corrections for sp3 carbon chemical shift calculations
The attached C carbons are:
C = -(2) + 9x(# + #) - 2x(# ) + steric corrections
The calculated
carbon atom is:
primary
quaternary
tertiary
primary
-1.1
-3.4
secondary
-2.5
-7.5
-15.0
tertiary
quaternary
13
secondary
-3.7
-9.5
-1.5
-8.4
-15.0
C1 = -2 + 9(1+3) - 2(2) + (-3) = 29
(actual = 28.3)
C2 = -2 + 9(4+2) - 2(2) + [3x(-1.5)+(-15.0)] = 28
2
C3 = -2 + 9(3+5) - 0(2) + [(-9.5)+(-15.0)] = 45
-25.0
(actual = 34.0)
(actual = 47.9)
C4 = -2 + 9(3+2) - 3(2) + (-9.5) = 27
C5 = -2 + 9(1+2) - 2(2) + (-1) = 20
(actual = 27.2)
(actual = 19.5)
C6 = -2 + 9(1+2) - 5(2) + (-1) = 14
(actual = 8.5)
C shifts for various carbon alkane skeletons - useful starting points for calculating sp3 carbon chemical shifts
C3
C2
CH4
-2.3
15.8
5.9
C6
22.9
14.1
28.1
48.9
29.3
18.8
39.0
33.9
29.8
29.7
34.4
39.2
35.3
14.5
25.2
C9
14.1
39.2
C10
22.8
29.6
32.1
29.8
26.9
42.3
14.2
29.6
22.8
27.2
28.1
40.6
38.0
22.7
32.1
12.0
31.8
17.9
37.2
C8
14.1
18.1
17.5
27.0
36.4
29.5
15.0
14.7
48.3
33.4
11.0
9.1
22.9
32.0
31.9
20.3
27.0
25.6
29.7
19.5
14.1
23.1
32.0
11.8
36.5
11.4
29.9
32.1
34.4
8.9
30.4
19.2
22.7
29.3
22.9
36.3
14.4
30.2
29.0
11.5
41.5
22.6
14.1
20.6
32.9
C7
25.0
13.8
27.9
22.3
C5
25.4
25.0
22.7
22.9
14.1
C4
16.3
32.3
14.1
22.9
29.6
32.2
Z:\files\classes\spectroscopy\typical spectra charts.DOC
11.5
34.6
14.5
36.5
19.3
29.7
14.1
20.3
39.6
32.4
19.7
29.9
10.9
40.3
25.6
35.4
20.1
14.6
Spectroscopy Data Tables
X
C correction C correction
C correction
X is attached to a terminal carbon atom (ppm)
Substituent = X
15
X is attached to an internal carbon atom (ppm)
C correction C correction
C correction
CH3
-2
-2
CH2CH3
18
-2
-2
CH(CH3)2
26
-2
14
-2
C(CH3)3
32
-2
20
-2
C
H
CH2
20
-1
15
-1
CH
-4
-4
23
-2
17
-2
X is attached to a terminal carbon atom (ppm)
Substituent = X
C correction C correction
C correction
X is attached to an internal carbon atom (ppm)
C correction C correction
C correction
OH
48
10
-6
44
-4
OR
60
-6
57
-6
51
-6
49
-6
NH2
28
10
-5
24
-5
NH(CH3)
38
-5
32
-4
N(CH3)2
45
-5
37
-4
-5
21
-5
-5
-5
O
O
C
R
O
H
N
26
C
R
NO2
62
Z:\files\classes\spectroscopy\typical spectra charts.DOC
58
Spectroscopy Data Tables
16
X is attached to a terminal carbon atom (ppm)
Substituent = X
C correction C correction
C correction
X is attached to an internal carbon atom (ppm)
C correction C correction
C correction
70
-7
67
-7
Cl
31
10
-5
36
-5
Br
20
10
-4
28
10
-4
-7
11
-2
11
-2
30
-3
24
-1
-3
31
-3
26
-3
22
-3
18
-3
O
C
H
O
C
CH3
O
C
OH
X is attached to a terminal carbon atom (ppm)
Substituent = X
C correction C correction
C correction
X is attached to an internal carbon atom (ppm)
C correction C correction
C correction
20
-3
16
-3
25
-3
19
-3
-3
-3
33
-3
30
-3
SH
11
10
-3
12
-3
SR
22
-3
20
-3
C
OCH3
O
C
NH2
C
N
O
C
Cl
Z:\files\classes\spectroscopy\typical spectra charts.DOC
Spectroscopy Data Tables
17
Additional starting point for calculating 13C chemical shifts (ppm) of substituted benzene rings (just a few possibilities)
Substituent
128 ppm starting point for
benzene carbon
Use correction term for carbon
atom in relative position to the
substituent. Start with 128 ppm.
Starting points for other common ring
systems. (ppm). No correction terms
included for substituents.
3
Z1
0
9
12
10
11
20
19
8
9
9
11
12
15
2
2
12
7
6
6
13
-6
8
34
5
-5
-31
Substituent
-H
-CH3
-CH2CH3
-CH2CH2CH3
-CH2CH2CH2CH3
-CH(CH3)2
-C(CH3)3
-CH2F
-CH2Cl
-CH2Br
-CH2I
-CH2OH
-CH2NH2
-CH2NO2
-CH2CN
-CH2SH
-CH2CHO
-CH2COCH3
-CH2CO2H
-CH2=CH2
-CCH
-C6H5
-F
-Cl
-Br
-I
Z2
0
1
-1
0
0
-2
-3
-1
0
1
-1
-1
-1
2
0
-1
1
1
1
-3
4
-1
-13
0
3
9
Z3
0
0
0
0
0
0
0
0
0
0
0
0
0
1
-1
0
0
0
0
0
0
0
2
1
2
2
Z4
0
-3
-3
-3
-3
-3
-3
0
0
0
-1
-1
-2
1
-1
-2
-1
-2
-1
-1
0
-1
-4
2
-1
-1
Z1
29
34
28
18
10
22
23
22
37
20
4
10
18
12
16
-16
8
9
2
2
5
11
-43
-36
Substituent
-OH
-OCH3
-OC6H5
-NH2
-NHCOCH3
-NHOH
-NHNH2
-N=N-R
-NO
-NO2
-SH
-SCH3
-S(O)CH3
-SO2CH3
-SO2Cl
-CN
-CHO
-COCH3
-CO2H
-CO2CH3
-CONH2
-COCl
-Li
-MgBr
Z2
-13
-14
-11
-13
-8
-13
-16
-6
-8
-5
1
-2
-5
-1
-2
3
1
0
2
1
-1
0
-13
-11
Z3
1
1
0
1
0
-2
1
0
1
1
0
0
1
1
1
1
0
0
0
0
0
0
2
3
126
128
Z4
-7
-8
-7
-10
-4
-5
-10
-3
7
6
-3
-4
2
5
7
4
6
4
5
4
3
-3
3
4
134
naphathalene
136
124
pyridine
150
108
pyrrole
N
H
118
110
furan
143
126
thiophene
125
Additional starting point for calculating 13C chemical shifts (ppm) of substituted alkenes (just a few possibilities)
123 ppm starting point for alkene carbon
'
C
'
C
Z
C
2
'
C
C = 123 ppm + Zi
123 + correction factors
increments for directly attached carbon atoms
' = -8
= 11
' = -2
=5
'= 2
= -2
steric corrections
for each pair of cis-, ' substituents
for each pair of geminal-, substituents
for each pair of geminal-, 'substituents
if one or more sutstituents are present
C
1
-1
-5
3
2
Z:\files\classes\spectroscopy\typical spectra charts.DOC
Effect of substituents on alkene 13C shifts (ppm)
Substituent
-H
-CH3
-CH2CH3
-CH2CH2CH3
-CH(CH3)2
-C(CH3)3
-CH2Cl
-CH2Br
-CH2I
-CH2OH
-CH=CH2
-CCH
-C6H5
Z1
0
13
17
16
23
26
10
11
14
14
14
-6
12
Z2
0
-7
-10
-9
-12
-15
-6
-5
-4
-8
-7
6
-11
Substituent
-F
-Cl
-Br
-I
-OCH3
-O2CCH3
-N(CH3 )2
-NO2
-CN
-SCH2CH3
-CHO
-COCH3
-CO2H
-COCl
Z1
24
3
-9
-38
29
18
28
22
-15
9
15
14
5
8
Z2
-34
-6
-1
7
-39
-27
-32
-1
14
-13
14
5
10
14
Spectroscopy Data Tables
18
Common fragmentation patterns in mass spectroscopy
1. Branch next to a bond
R
C
radical cation
Pi electrons partially fill in loss of electrons at
carbocation site via resonance. This is common
fragmentation for alkenes and aromatics
bond of an alkene
or an aromatic
Characteristic carboncation stability also applies.
3o R
>
2o R
> 1o R
>
CH3
2. Branch next to an atom with a lone pair of electrons
R
X C
X C
X C
radical cation
X lone pair electrons partially fill in loss of electrons at carbocation site via resonance. This
is a common fragmentation for any atom that has a lone pair of electrons (oxygen = alcohol,
ether, ester; nitrogen = amine, amide, sulfur = thiol or sulfide, etc.). Alcohols often lose water
(M-18) and primary amines can lose ammonia (M-17).
3. Branch next to a carbonyl (C=O) bondand possible subsequent loss of carbon monoxide, CO
R1
O
R1
C O
R1
R1
C O
loss of
R2
R2
radical cation
R1 or R2 can be lost from
aldehydes, ketones, acids,
esters, amides...etc.
R2
C O
C O
An oxygen lone pair partially fill in the loss of electrons at the
carbocation site via resonance. This is a common fragmentation
pattern for any carbonyl compound and can occur from either
side, though some are more common than others.
R2
subsequent loss of CO is possible
after fragmentation so not only can
you see loss of an branch you can
see the mass of an branch.
4. McLafferty Rearrangement
O
R1
H
C
C
C
O
R1
H
C
C
C
Positive charge can be on
either fragment, which
typically have an even mass.
= alpha position
= beta position
= gama position
lost neutral
still a radical
fragment
cation
This is another common fragmentation pattern for carbonyl compounds (and other pi systems as well: alkenes, aromatics,
alkynes, nitriles, etc.). If the pi bond has at least 3 additional nonhydrogen atoms attached and a hydrogen on the "gama"
atom, the branch can curve around to a comfortable 6 atom arrangement and the pi bond can pick up a hydrogen atom and
cut off a fragment between the C and C positions. The positive charge can be seen on either fragment and usually the
fragments have an even mass (unless there is an odd number of nitrogen atoms).
radical cation
Knowing these few fragmentation patterns will allow you to make many useful predictions and
interpretations. Loss of small molecules, via elimination is common: H2O = 18, H2S = 34, CH3OH = 32,
C2H5OH = 46, NH3 = 17, CH3CO2H = 62, HF = 20, HCl = 36/38, HBr = 80/82, etc.
Z:\files\classes\spectroscopy\typical spectra charts.DOC
Spectroscopy Data Tables
19
A sampling of unusual and/or miscellaneous peaks that are commonly seen, (even when they don't make sense).
R
CH3 = 15
CH3CH2 = 29
C3H7 = 43
C4H9 = 57
C5H11 = 71
C6H13 = 85
mass = 39 (R = H)
53 (R = CH3)
67 (R= CH2CH3)
also works for
R
CH2
C
H
mass = 41 (R = H)
55 (R = CH3)
69 (R= CH2CH3)
H2N
mass = 65 (R = H)
79 (R = CH3)
93 (R= CH2CH3)
mass = 27
mass = 77
RO
mass = 91 (R = H)
105 (R = CH3)
119 (R= CH2CH3)
C
H2
mass = 29 (R = H)
43 (R = CH3)
mass = 42 (R = H)
57 (R= CH2CH3)
56 (R = CH3)
71 (R = C3H7)
70 (R= CH2CH3)
105 (R = C6H5)
mass = 44
H
mass = 45 (R = H)
59 (R = CH3)
73 (R= CH2CH3)
Loss of small molecules via elimination reactions.
H2 O
mass = 18
CH3OH
32
H2 S
34
C2H5OH
46
HF HCl HBr
80
20 36 82
38
NH3 CH3CO2H
62
17
McLafferty Possibilities
H
H
O
O
R2
HC
R1
CH2
C
H2
R2
R1
R
McLafferty
Notice!
even masses
mass = 44 (R = H)
58 (R = CH3)
72 (R= CH2CH3)
86 (R = C3H7)
CH2
variable mass,
(can sometimes see
cation on this side too)
mass = 28 (R = H)
42 (R = CH3)
56 (R= CH2CH3)
70 (R = C3H7)
Similar Patterns
H
CH2
R2
R1
H
R2
H
H
R1
mass = 42 (R = H)
56 (R = CH3)
70 (R= CH2CH3)
84 (R = C3H7)
R1
mass = 92 (R = H)
106 (R = CH3)
120 (R= CH2CH3)
134 (R = C3H7)
Z:\files\classes\spectroscopy\typical spectra charts.DOC
R2
R1
C
H2
H
C
R2
R1
CH2
C
H2
R2
R
R2
R1
C
H2
H
H2C
R1
C
H2
R2
C
CH2
mass = 40 (R = H)
54 (R = CH3)
68 (R= CH2CH3)
82 (R = C3H7)
R2
R1
CH2
mass = 41 (R = H)
55 (R = CH3)
69 (R= CH2CH3)
83 (R = C3H7)
You might also like
- JBL Audio Engineering for Sound ReinforcementFrom EverandJBL Audio Engineering for Sound ReinforcementRating: 5 out of 5 stars5/5 (2)
- HC-Bàn Luận Hữu Cơ-Nguyễn Văn PhòngDocument393 pagesHC-Bàn Luận Hữu Cơ-Nguyễn Văn PhòngDat Vu100% (1)
- Schaum's Easy Outline of Organic Chemistry, Second EditionFrom EverandSchaum's Easy Outline of Organic Chemistry, Second EditionRating: 3.5 out of 5 stars3.5/5 (2)
- Notes The Common and Iupac Names of Organic CompoundsDocument2 pagesNotes The Common and Iupac Names of Organic Compoundszaibakhan817% (6)
- Spec Ir NMR Spectra TablesDocument15 pagesSpec Ir NMR Spectra TablesMah NovaesNo ratings yet
- Infrared (IR) Spectroscopy: Structure, Purity, and IdentityDocument16 pagesInfrared (IR) Spectroscopy: Structure, Purity, and IdentityDiana KowsariNo ratings yet
- IR ProcedureDocument5 pagesIR ProcedurePuvaneswary LoganathanNo ratings yet
- Functional Class Range (NM) Intensity Assignment Range (NM) Intensity AssignmentDocument6 pagesFunctional Class Range (NM) Intensity Assignment Range (NM) Intensity AssignmentdubstepoNo ratings yet
- IR ProcedureDocument5 pagesIR ProcedureMuhammad FauziNo ratings yet
- Infrared Spectroscopy: Conformational IsomersDocument7 pagesInfrared Spectroscopy: Conformational IsomersRiyan NazarudinNo ratings yet
- Spektro IRDocument64 pagesSpektro IRAnonymous NSK4nvH4ufNo ratings yet
- 2230L 08 IR Spectra InterpretationDocument11 pages2230L 08 IR Spectra Interpretationvennilaj23No ratings yet
- ASS Instrumental OrganicDocument17 pagesASS Instrumental OrganicMohamed SakrNo ratings yet
- Infrared Spectroscopy: Concepts and TheoriesDocument55 pagesInfrared Spectroscopy: Concepts and Theoriesdead_knightNo ratings yet
- Lecture 4 IR Spectrum AnalysisDocument43 pagesLecture 4 IR Spectrum AnalysiskhadijahhannahNo ratings yet
- IR-freq CO BondDocument3 pagesIR-freq CO BondRD's AcademyNo ratings yet
- Raw Material Analysis-IRDocument58 pagesRaw Material Analysis-IRDilla Wulan NingrumNo ratings yet
- 08 - Infrared Spectroscopy ManualDocument5 pages08 - Infrared Spectroscopy ManualShubham BobadeNo ratings yet
- Infrared Correlations: Functional Group Band Position (CM) AppearanceDocument2 pagesInfrared Correlations: Functional Group Band Position (CM) AppearanceAmritansh RanjanNo ratings yet
- Key HW 3 Part II SpecDocument16 pagesKey HW 3 Part II SpecTha KantanaNo ratings yet
- PHR410 Chapter 2Document36 pagesPHR410 Chapter 2pulock.paulNo ratings yet
- Ir Func GroupDocument52 pagesIr Func GroupEry NourikaNo ratings yet
- 6-IR Spectroscopy of Alkane, Alkene and Carbonyl CompoundsDocument8 pages6-IR Spectroscopy of Alkane, Alkene and Carbonyl Compoundsbloodhound13042005No ratings yet
- Spec Ir NMR Spectra Tables PDFDocument15 pagesSpec Ir NMR Spectra Tables PDFYuppie RajNo ratings yet
- Introduction To Interpretation of Infrared SpectraDocument3 pagesIntroduction To Interpretation of Infrared SpectraBenni WewokNo ratings yet
- IR SPECTROSCOPY Notes FullDocument5 pagesIR SPECTROSCOPY Notes FullKartik KuteNo ratings yet
- Spectroscopy Infrared SpectraDocument51 pagesSpectroscopy Infrared Spectrathanasa08No ratings yet
- SPECTRA TablesDocument19 pagesSPECTRA TablesMiroslav VetrikNo ratings yet
- FTIR TablesDocument1 pageFTIR TablesvandykavidurgaNo ratings yet
- IR SpectrosDocument33 pagesIR SpectrosKikiMariaNo ratings yet
- IR Spectra AnalysisDocument37 pagesIR Spectra AnalysisdevoydouglasNo ratings yet
- Introduction To Interpretation of Infrared SpectraDocument3 pagesIntroduction To Interpretation of Infrared Spectrachinnirao100% (4)
- Topic 9 NotesDocument9 pagesTopic 9 NotesRitik YadavNo ratings yet
- IRSpectrum AnalysisDocument2 pagesIRSpectrum AnalysisDavid S. FrohnapfelNo ratings yet
- Common I R Absorption SDocument1 pageCommon I R Absorption SVisakha SureshNo ratings yet
- IR Spectroscopy TutorialDocument36 pagesIR Spectroscopy TutorialreddygrNo ratings yet
- Functional Groups Functional Groups: Functional Group G PDocument52 pagesFunctional Groups Functional Groups: Functional Group G PZenonissya Galwan BataraNo ratings yet
- Interpretation of Spectra of Different CompoundsDocument15 pagesInterpretation of Spectra of Different Compoundsmariam nawabNo ratings yet
- Experiment 2 Laboratory Manual 2022Document13 pagesExperiment 2 Laboratory Manual 2022Nicoleta MaritanuNo ratings yet
- Spec IR Table For Common Chemical SymbolsDocument4 pagesSpec IR Table For Common Chemical SymbolsYoussef LatashNo ratings yet
- Printable Acrobat PDF File: Table of Characteristic IR AbsorptionsDocument3 pagesPrintable Acrobat PDF File: Table of Characteristic IR AbsorptionsImam Hadillah MuhfiNo ratings yet
- Table - 1: Characteristic Infrared Absorptions of Functional GroupsDocument1 pageTable - 1: Characteristic Infrared Absorptions of Functional GroupsAJIT CHAUDHARINo ratings yet
- CHMBD 449 - Organic Spectral: AnalysisDocument40 pagesCHMBD 449 - Organic Spectral: AnalysisIleana ManciuleaNo ratings yet
- Infrared SpectrosDocument110 pagesInfrared SpectrosBHARTI GAURNo ratings yet
- Study Notes-IR SpectrosDocument24 pagesStudy Notes-IR SpectrosakshantratwanNo ratings yet
- Ir PDFDocument1 pageIr PDFBartłomiej LesiszNo ratings yet
- CHMBD 449 - Organic Spectral: AnalysisDocument43 pagesCHMBD 449 - Organic Spectral: AnalysisIleana ManciuleaNo ratings yet
- Infrared Tutorial 2Document71 pagesInfrared Tutorial 2Hammo Ez AldienNo ratings yet
- IR ChartDocument2 pagesIR ChartNadiaa SafirraNo ratings yet
- Solomons Organic Chemistry Module IR TableDocument1 pageSolomons Organic Chemistry Module IR TableBenni WewokNo ratings yet
- Functional Class Range (CM) Intensity Assignment Alkanes: AlkenesDocument1 pageFunctional Class Range (CM) Intensity Assignment Alkanes: AlkenesStoica AlexandruNo ratings yet
- Spektrometri IRDocument51 pagesSpektrometri IRClarion 642No ratings yet
- FullDocument10 pagesFullAbdul Wahab KhanNo ratings yet
- IR SpectrosDocument44 pagesIR SpectrosVansh YadavNo ratings yet
- Alcohol: Functional Group Type of Vibration Characteristic Absorptions (CM) IntensityDocument2 pagesAlcohol: Functional Group Type of Vibration Characteristic Absorptions (CM) IntensityMuhammad Fadhila Ragil YogaNo ratings yet
- Scanning Electron Microscopy (SEM) With Energy Dispersive Spectroscopy (EDS) AnalysisDocument5 pagesScanning Electron Microscopy (SEM) With Energy Dispersive Spectroscopy (EDS) AnalysisAjeeth KumarNo ratings yet
- Infrared Spectroscopy: IR Absorptions For Representative Functional GroupsDocument3 pagesInfrared Spectroscopy: IR Absorptions For Representative Functional GroupsSaleem BashaNo ratings yet
- Measures Molecular Vibrations of Characteristic Functional GroupsDocument4 pagesMeasures Molecular Vibrations of Characteristic Functional GroupsLejNo ratings yet
- Vibrational Spectra of Organometallics: Theoretical and Experimental DataFrom EverandVibrational Spectra of Organometallics: Theoretical and Experimental DataNo ratings yet
- Images from Lichenes Australasici Exsiccati and of other characteristic Australasian Lichens. Volume OneFrom EverandImages from Lichenes Australasici Exsiccati and of other characteristic Australasian Lichens. Volume OneNo ratings yet
- Discrete Series of GLn Over a Finite Field. (AM-81), Volume 81From EverandDiscrete Series of GLn Over a Finite Field. (AM-81), Volume 81No ratings yet
- Bảng phổ IRDocument5 pagesBảng phổ IRĐan KhanhNo ratings yet
- Solutions AIATS JEE (Main) - 2017 Test-7 Paper-1 (Code-A & B) (19!02!2017)Document20 pagesSolutions AIATS JEE (Main) - 2017 Test-7 Paper-1 (Code-A & B) (19!02!2017)Jalaj LabanaNo ratings yet
- C) Trigonal Planar: E-Pent-2-ene Z-Pent-2-ene Z-3-Methylpent-2-ene Z-2-Methylpent-2-eneDocument9 pagesC) Trigonal Planar: E-Pent-2-ene Z-Pent-2-ene Z-3-Methylpent-2-ene Z-2-Methylpent-2-eneJessicaNo ratings yet
- 6carboxylic AcidsDocument1 page6carboxylic AcidssharmimiameerasanadyNo ratings yet
- AminesDocument23 pagesAminesfhtzzzzzzNo ratings yet
- CHAPTER 8 EditedDocument18 pagesCHAPTER 8 EditedSyafiqah SuhaimiNo ratings yet
- CarbenesDocument4 pagesCarbenesDr_GSNo ratings yet
- Aldehyde Ketones and Carboxylic AcidDocument18 pagesAldehyde Ketones and Carboxylic AcidInfinite SinghNo ratings yet
- Chapter 19Document26 pagesChapter 19tyobertsNo ratings yet
- Organic Compounds Containing OxygenDocument17 pagesOrganic Compounds Containing OxygenSai PrajitNo ratings yet
- Reaction With Miscellaneous-NPTEL PDFDocument25 pagesReaction With Miscellaneous-NPTEL PDFRathinNo ratings yet
- HSPi PData SetDocument210 pagesHSPi PData Setkamilo14100% (1)
- Important Reactions For Iit JeeDocument4 pagesImportant Reactions For Iit JeeRajesh RanjanNo ratings yet
- c7h15 Oh Ch3 Ch2 Ch2 Ch2 Ch2 Ch2 Ch2 Oh Oh Ch3 Ch2 Ch2 Ch2 Ch2 Ch2 Ch3 Oh Ch3 Ch2 Ch2 Ch2 Ch2 Ch2 Ch3 OhDocument15 pagesc7h15 Oh Ch3 Ch2 Ch2 Ch2 Ch2 Ch2 Ch2 Oh Oh Ch3 Ch2 Ch2 Ch2 Ch2 Ch2 Ch3 Oh Ch3 Ch2 Ch2 Ch2 Ch2 Ch2 Ch3 OhrizqieNo ratings yet
- HydrocarbonsDocument37 pagesHydrocarbonsraghavsuresh865No ratings yet
- 8 Esters Have Many Uses Due To Their Characteristic Aromas and Often Have CommonDocument3 pages8 Esters Have Many Uses Due To Their Characteristic Aromas and Often Have CommonMohamed MuhajireenNo ratings yet
- 12th Board Sprint-Amines (15.12.2020)Document62 pages12th Board Sprint-Amines (15.12.2020)Harsh ShahNo ratings yet
- 16H Carbonyl PDFDocument60 pages16H Carbonyl PDFJose Erick Ortega ValenciaNo ratings yet
- DPPONIUPACSUPERSIXER4Document5 pagesDPPONIUPACSUPERSIXER4Kartik YadavNo ratings yet
- Chapter 21. Carboxylic Acid Derivatives: Nucleophilic Acyl Substitution ReactionsDocument20 pagesChapter 21. Carboxylic Acid Derivatives: Nucleophilic Acyl Substitution Reactions張湧浩No ratings yet
- Complete Course in Organic Chemistry by Vineet Khatri Sir: Class: Xi Time: 35 Min. DPP. NO.17Document4 pagesComplete Course in Organic Chemistry by Vineet Khatri Sir: Class: Xi Time: 35 Min. DPP. NO.17Arnab KumarNo ratings yet
- OrganicDocument3 pagesOrganicSchimmel Repelin Velud Aytresogres100% (1)
- Mock Exam 2-AnswersDocument8 pagesMock Exam 2-AnswersKhaledEl-MaghallawyNo ratings yet
- Carboxylic Acid and Amines Worksheet PDFDocument22 pagesCarboxylic Acid and Amines Worksheet PDFd anjilappaNo ratings yet
- Aldehyde, Ketone and Carboxylic Acid Class 12 CbseDocument8 pagesAldehyde, Ketone and Carboxylic Acid Class 12 CbseRahul SharmaNo ratings yet
- Annex 10 Ordinance Fdha Materials and Articles Intended To Come Into Contact With Food StuffsDocument202 pagesAnnex 10 Ordinance Fdha Materials and Articles Intended To Come Into Contact With Food Stuffsjai soniNo ratings yet
- 9 - H Eterofunctional CONNECTIONS - Aliphatic and Benzene SeriesDocument10 pages9 - H Eterofunctional CONNECTIONS - Aliphatic and Benzene SeriesAshish SingrohaNo ratings yet
- كيمياء حيوية الوحدة التانيةDocument50 pagesكيمياء حيوية الوحدة التانيةasem sawalmehNo ratings yet