Professional Documents
Culture Documents
MP Hpohiperglikemia
MP Hpohiperglikemia
Uploaded by
Yanuar PrasetyoCopyright:
Available Formats
You might also like
- 2022 Book ProtocolsForTheDiagnosisOfPigVDocument383 pages2022 Book ProtocolsForTheDiagnosisOfPigV谭硕100% (1)
- Textbook of Practical MicrobiologyDocument329 pagesTextbook of Practical MicrobiologyMara Maruca94% (50)
- Cytokine Bioassays PDFDocument383 pagesCytokine Bioassays PDFSubash Suba100% (1)
- 2 PMID 37992081 BiofilmDocument15 pages2 PMID 37992081 Biofilmajaya basnetNo ratings yet
- 2Document12 pages2Annu OjhaNo ratings yet
- Mycoplasma Pneumoniae: Prevalance in Navi Mumbai, India Biomedical. Indian Journal of Applied ResearchDocument4 pagesMycoplasma Pneumoniae: Prevalance in Navi Mumbai, India Biomedical. Indian Journal of Applied ResearchSpinalis 2017No ratings yet
- Tube Method and Congo Red Agar Versus Tissue CultuDocument7 pagesTube Method and Congo Red Agar Versus Tissue CultuMohanad TofahNo ratings yet
- Microbiological Laboratory Techniques - Clinical GateDocument18 pagesMicrobiological Laboratory Techniques - Clinical GateCamila Carla CapriglioniNo ratings yet
- Study of Extended Spectrum Beta Lactamases & Biofilm in Pseudomonas Species Isolated From Different Clinical SamplesDocument7 pagesStudy of Extended Spectrum Beta Lactamases & Biofilm in Pseudomonas Species Isolated From Different Clinical SamplesIJAR JOURNALNo ratings yet
- Prospective Multicentre Evaluation of The Direct NDocument5 pagesProspective Multicentre Evaluation of The Direct Npooja pandeyNo ratings yet
- BMS Iddt 2020 41Document8 pagesBMS Iddt 2020 41Guneyden GuneydenNo ratings yet
- 2022 Article 19844Document12 pages2022 Article 19844raphaNo ratings yet
- CV Abdul HafeezDocument7 pagesCV Abdul HafeezAbdul Hafeez KandhroNo ratings yet
- Diagnosing: Jericko MagistradoDocument25 pagesDiagnosing: Jericko MagistradoJericko MagistradoNo ratings yet
- Periodic Acid Schiff Reactions and General Tissue Morphology of Conventionally Processed Versus Two Rapid Microwave Processed TissuesDocument15 pagesPeriodic Acid Schiff Reactions and General Tissue Morphology of Conventionally Processed Versus Two Rapid Microwave Processed TissuesRestiyaniNo ratings yet
- HHS Public Access: Getting Started With Microbiome Analysis: Sample Acquisition To BioinformaticsDocument41 pagesHHS Public Access: Getting Started With Microbiome Analysis: Sample Acquisition To Bioinformatics@lsreshtyNo ratings yet
- Basic Microbiological TestsDocument13 pagesBasic Microbiological TestsyeabsraNo ratings yet
- Guidelinesfor Quick Applicationof Biochemical Teststo Identify Unknown BacteriaDocument19 pagesGuidelinesfor Quick Applicationof Biochemical Teststo Identify Unknown Bacteriara.rodriguezojedaNo ratings yet
- Microbiological Examination For Suspected BacteriaDocument8 pagesMicrobiological Examination For Suspected BacteriaAhmad Alfiansyah SaidNo ratings yet
- Journal of Biosafety and BiosecurityDocument5 pagesJournal of Biosafety and BiosecuritypratiwiNo ratings yet
- Antibio AbstractDocument2 pagesAntibio Abstractvaidyam78No ratings yet
- Laboratory Guide Methodologies For Antimicrobial Susceptibility TestingDocument19 pagesLaboratory Guide Methodologies For Antimicrobial Susceptibility TestingSharina Mae Quilaquiga FidelNo ratings yet
- Mag Hale MostafaDocument8 pagesMag Hale MostafaFranco SantinNo ratings yet
- First Priniciples of Clinical MicrobiologyDocument7 pagesFirst Priniciples of Clinical MicrobiologyDi MonizNo ratings yet
- 45 Supplement - 2 S99Document13 pages45 Supplement - 2 S99Mustaqeem DawarNo ratings yet
- Cells 10 00463 v2Document40 pagesCells 10 00463 v2Stephanie Nichole Ian CasemNo ratings yet
- Formulation and Evaluation of Acyclovir MicrocapsuDocument5 pagesFormulation and Evaluation of Acyclovir MicrocapsuRizQi FatmiyahNo ratings yet
- Principle and Techniques of Immunohistochemistry A ReviewDocument8 pagesPrinciple and Techniques of Immunohistochemistry A ReviewAyunda Novita RaniNo ratings yet
- Biocompatibility Test Methods - Pacific BioLabsDocument11 pagesBiocompatibility Test Methods - Pacific BioLabsAnil KumarNo ratings yet
- Coagulase Negative Staphylococcus Species : Resistance and Therapeutic Decisions at The Turn of The Novel MillenniumDocument6 pagesCoagulase Negative Staphylococcus Species : Resistance and Therapeutic Decisions at The Turn of The Novel MillenniumdrpklalNo ratings yet
- Maldi-Tof MsDocument12 pagesMaldi-Tof MsD. GandhirajNo ratings yet
- Association of Virulence Genes With Antibiotic Resistance in Pakistani Uropathogenic E. Coli IsolatesDocument6 pagesAssociation of Virulence Genes With Antibiotic Resistance in Pakistani Uropathogenic E. Coli Isolateskhurry testNo ratings yet
- A Practical Guide To ISO 10993-5 - Cytotoxicity - MDDI Medical Device and Diagnostic Industry News Products and SuppliersDocument4 pagesA Practical Guide To ISO 10993-5 - Cytotoxicity - MDDI Medical Device and Diagnostic Industry News Products and SuppliersVanessa DuzNo ratings yet
- FTIR Analysis of Natural and Synthetic Collagen: Applied Spectroscopy Reviews March 2018Document46 pagesFTIR Analysis of Natural and Synthetic Collagen: Applied Spectroscopy Reviews March 2018Imelda Irma IsauraNo ratings yet
- Jomb 2014 0043 PDFDocument9 pagesJomb 2014 0043 PDFGonzalez ArturoNo ratings yet
- B.sc. Medical Laboratory Sciences Medical Bacteriology (MLS 2413) (PDFDrive)Document101 pagesB.sc. Medical Laboratory Sciences Medical Bacteriology (MLS 2413) (PDFDrive)Isah MohammedNo ratings yet
- 12 Catheter TipDocument6 pages12 Catheter Tipselvakumar selvarajNo ratings yet
- 404-Article Text-2908-1-10-20230508Document14 pages404-Article Text-2908-1-10-20230508elbizco8No ratings yet
- An Assignment On Methods of Sterilization: Submitted ToDocument35 pagesAn Assignment On Methods of Sterilization: Submitted ToShahin Rahman 112700gmail.comNo ratings yet
- Detection of Antibiotic Resistance and Bio Film-Producing Ability of Staphylococcus Species in Clinical IsolatesDocument5 pagesDetection of Antibiotic Resistance and Bio Film-Producing Ability of Staphylococcus Species in Clinical Isolatesafnan jumaaNo ratings yet
- The Term Antibiotic Refers To Any Substance Produced by Any Microbial Entity That Is Detrimental To Other Other Microbial SpeciesDocument26 pagesThe Term Antibiotic Refers To Any Substance Produced by Any Microbial Entity That Is Detrimental To Other Other Microbial SpeciesPratibha PatilNo ratings yet
- Biosafety in Microbiological Laboratories M A LaysiasingaporeindiaDocument385 pagesBiosafety in Microbiological Laboratories M A Laysiasingaporeindiamohd farihan musthafa kamalNo ratings yet
- Anticancer Activities of Antimicrobial Bmkn2 Peptides Against Oral and Colon Cancer CellsDocument12 pagesAnticancer Activities of Antimicrobial Bmkn2 Peptides Against Oral and Colon Cancer CellselustosacamposNo ratings yet
- 1 s2.0 S2225411022000554 MainDocument11 pages1 s2.0 S2225411022000554 MaindanielwongtsNo ratings yet
- 30395-Article Text-56938-1-10-20220916Document8 pages30395-Article Text-56938-1-10-20220916Djim KARYOMNo ratings yet
- Evaluation of Diffusion and Dilution Methods To Determine The Antibacterial Activity of Plant ExtractsDocument7 pagesEvaluation of Diffusion and Dilution Methods To Determine The Antibacterial Activity of Plant ExtractsHusniya FaradisaNo ratings yet
- Wa0011.Document11 pagesWa0011.umayamajidNo ratings yet
- Integrating Molecular Biology Into The Veterinary CurriculumDocument17 pagesIntegrating Molecular Biology Into The Veterinary CurriculumSin PermahaNo ratings yet
- Dont KnowDocument5 pagesDont KnownawarajNo ratings yet
- PCR Optimization For Beginners A Step by Step GuideDocument23 pagesPCR Optimization For Beginners A Step by Step GuideĐặng Gia HoàngNo ratings yet
- Paso Corto 7 - Diagnostic Microbiology in Veterinary Dermatology-1Document10 pagesPaso Corto 7 - Diagnostic Microbiology in Veterinary Dermatology-1ABIGAIL AKEMI RODRIGUEZ CHAVEZNo ratings yet
- Guidance On Good Cell Culture PracticeDocument28 pagesGuidance On Good Cell Culture PracticeNyna KawlesNo ratings yet
- Diagnosis of Bacterial Infections (QH 29.02.2016)Document50 pagesDiagnosis of Bacterial Infections (QH 29.02.2016)rasool ghaffariNo ratings yet
- Screening and Selection of Industrially Important Microorganisms: A ReviewDocument7 pagesScreening and Selection of Industrially Important Microorganisms: A Reviewjeevalakshmanan29No ratings yet
- 1 s2.0 S1021949819300146 MainDocument11 pages1 s2.0 S1021949819300146 MainraraNo ratings yet
- BacterialisolationDocument10 pagesBacterialisolationMark SudieNo ratings yet
- Reviews:: Protein Cancer Biomarkers Serve MultiDocument11 pagesReviews:: Protein Cancer Biomarkers Serve MultiIoana CreangaNo ratings yet
- Comparison of Chemiluminescence Enzyme ImmunoassayDocument7 pagesComparison of Chemiluminescence Enzyme ImmunoassayDewi ArisandyNo ratings yet
- Diagnosis MikrobiologiDocument14 pagesDiagnosis Mikrobiologiarmelya aisyahNo ratings yet
- Analytical MicrobiologyFrom EverandAnalytical MicrobiologyFrederick KavanaghNo ratings yet
- WerwerwerDocument1 pageWerwerwerYanuar PrasetyoNo ratings yet
- DasdadasdDocument4 pagesDasdadasdYanuar PrasetyoNo ratings yet
- E.G. OP-11-Cherry - Pptx. For Your Presentation Code, Please Refer To The List of OralDocument2 pagesE.G. OP-11-Cherry - Pptx. For Your Presentation Code, Please Refer To The List of OralYanuar PrasetyoNo ratings yet
- Lai Et Al-2013-Arthritis & RheumatismDocument1 pageLai Et Al-2013-Arthritis & RheumatismYanuar PrasetyoNo ratings yet
- List of Oral Presenters: Time (WIB) Presentation Code Name RoomDocument2 pagesList of Oral Presenters: Time (WIB) Presentation Code Name RoomYanuar PrasetyoNo ratings yet
- Inhibitor-Based Methods For Detection of Plasmid-Mediated Ampc - Lactamases in Klebsiella SPP., Escherichia Coli, and Proteus MirabilisDocument5 pagesInhibitor-Based Methods For Detection of Plasmid-Mediated Ampc - Lactamases in Klebsiella SPP., Escherichia Coli, and Proteus MirabilisYanuar PrasetyoNo ratings yet
- Rounds: Case Study and Review of Autoimmune HepatitisDocument7 pagesRounds: Case Study and Review of Autoimmune HepatitisClaudia Naomi Ventura OrtizNo ratings yet
- Human Fertilization Is The Union of A Human Egg and SpermDocument7 pagesHuman Fertilization Is The Union of A Human Egg and Spermoxidalaj100% (1)
- Stem Cell Powerpoint With Teacher NotesDocument36 pagesStem Cell Powerpoint With Teacher NotesDELLA BLATAMANo ratings yet
- Influence of Antiretroviral Therapy On Maternal Eosinophil Levels During Pregnancy: A ReviewDocument14 pagesInfluence of Antiretroviral Therapy On Maternal Eosinophil Levels During Pregnancy: A ReviewKIU PUBLICATION AND EXTENSIONNo ratings yet
- Department of Internal Medicine: Non-Hodgkin LymphomaDocument26 pagesDepartment of Internal Medicine: Non-Hodgkin LymphomaHana FauziNo ratings yet
- Culture Media Culture Media PurposeDocument2 pagesCulture Media Culture Media Purposeangelique wongNo ratings yet
- The Complete Blood Cell Count A PowerfulDocument16 pagesThe Complete Blood Cell Count A PowerfulGabriele VitorNo ratings yet
- Repurposing Non Oncology Small Molecule Drugs To Improve - 2022 - Acta PharmaceDocument26 pagesRepurposing Non Oncology Small Molecule Drugs To Improve - 2022 - Acta PharmaceMohammed Shuaib AhmedNo ratings yet
- Rennets: MicrobialDocument35 pagesRennets: MicrobialJonathan CisternasNo ratings yet
- Fosbac Plus T (F+T) : in Broiler ProductionDocument25 pagesFosbac Plus T (F+T) : in Broiler Productionthanh ba matNo ratings yet
- The Histone Code HypothesisDocument5 pagesThe Histone Code HypothesisBrian D. StrahlNo ratings yet
- MicrosatellitesDocument24 pagesMicrosatellitesAhmed BioNo ratings yet
- Salmonella Pathogenicity Islands Encoding Type III Secretion SystemsDocument11 pagesSalmonella Pathogenicity Islands Encoding Type III Secretion SystemsIsabella GriffithNo ratings yet
- Introduction To Cell BiologyDocument43 pagesIntroduction To Cell BiologyEllemrac GageloniaNo ratings yet
- HAP Lab Quiz (Midterm)Document11 pagesHAP Lab Quiz (Midterm)shane.erica.penitNo ratings yet
- Evaluation of The Performance of Sysmex XN-3100 Automated Hematology Analyzer Regarding The Sysmex XE-2100 and Microscopic ExaminationDocument9 pagesEvaluation of The Performance of Sysmex XN-3100 Automated Hematology Analyzer Regarding The Sysmex XE-2100 and Microscopic ExaminationbalkisNo ratings yet
- Culture Media: For Industrial MicrobiologyDocument34 pagesCulture Media: For Industrial MicrobiologyKATHENo ratings yet
- What Are The HLA Genes?: Humans Have Three Main MHC Class I Genes, Known As HLA-A, HLA-B, and HLA-C. TheDocument2 pagesWhat Are The HLA Genes?: Humans Have Three Main MHC Class I Genes, Known As HLA-A, HLA-B, and HLA-C. ThepoojaNo ratings yet
- Monoclonal AntibodiesDocument35 pagesMonoclonal Antibodiespreetylyall100% (5)
- Mark Blood BankDocument20 pagesMark Blood BankMark Lester MoralesNo ratings yet
- Review Sheet On 7.2Document9 pagesReview Sheet On 7.2KJannat Mahmood owaitanNo ratings yet
- QUALITY MANUAL v008 FinalDocument22 pagesQUALITY MANUAL v008 FinalPERICODELOSPALOTESNo ratings yet
- Ingles Area1 Jan2016Document4 pagesIngles Area1 Jan2016Eugênio RegoNo ratings yet
- Litareture ReviewDocument3 pagesLitareture ReviewShowkatul IslamNo ratings yet
- Dr.R.a.reizkhi TugasDocument13 pagesDr.R.a.reizkhi TugasReizkhi FitriyanaNo ratings yet
- Principles of ImmunizationDocument4 pagesPrinciples of ImmunizationDoc Prince CaballeroNo ratings yet
- WES WGS Brochure Pages V2.1eng Web 20201019Document16 pagesWES WGS Brochure Pages V2.1eng Web 20201019drumerNo ratings yet
- Booster Mock Test For NEETDocument15 pagesBooster Mock Test For NEETDrNo ratings yet
- Virology Respiratory VirusesDocument6 pagesVirology Respiratory VirusesHisham ChomanyNo ratings yet
MP Hpohiperglikemia
MP Hpohiperglikemia
Uploaded by
Yanuar PrasetyoOriginal Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
MP Hpohiperglikemia
MP Hpohiperglikemia
Uploaded by
Yanuar PrasetyoCopyright:
Available Formats
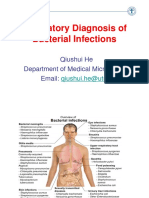
See discussions, stats, and author profiles for this publication at: https://www.researchgate.
net/publication/51589703
Evaluation of different detection methods of biofilm formation in the clinical
isolates
Article in The Brazilian journal of infectious diseases: an official publication of the Brazilian Society of Infectious Diseases · August 2011
DOI: 10.1016/S1413-8670(11)70197-0 · Source: PubMed
CITATIONS READS
94 1,412
6 authors, including:
Javaid Usman Fatima Kaleem
Army Medical College Foundation University Medical College, Islamabad, Pakistan
55 PUBLICATIONS 349 CITATIONS 46 PUBLICATIONS 336 CITATIONS
SEE PROFILE SEE PROFILE
Ali Khalid Muhammad Iqbal
King Abdullah Medical Complex Tanjungpura University
28 PUBLICATIONS 307 CITATIONS 25 PUBLICATIONS 250 CITATIONS
SEE PROFILE SEE PROFILE
Some of the authors of this publication are also working on these related projects:
Professor of Pathology (Microbiology), clinical microbiologist, researcher View project
All content following this page was uploaded by Ali Khalid on 21 May 2014.
The user has requested enhancement of the downloaded file.
ORIGINAL
ARTICLE
Evaluation of different detection methods
of biofilm formation in the clinical isolates
ABSTRACT Authors
Afreenish Hassan1
Background: Microorganisms growing in a biofilm are associated with chronic and recurrent hu- Javaid Usman2
man infections and are highly resistant to antimicrobial agents. There are various methods to detect Fatima Kaleem1
Maria Omair3
biofilm production like Tissue Culture Plate (TCP), Tube method (TM), Congo Red Agar meth- Ali Khalid1
od (CRA), bioluminescent assay, piezoelectric sensors, and fluorescent microscopic examination. Muhammad Iqbal4
Objective: This study was conducted to compare three methods for the detection of biofilms.
MPhil Microbiology,
Method: The study was carried out at the Department of Microbiology, Army Medical College,
1
Department of Microbiology,
National University of Sciences and Technology, Pakistan, from January 2010 to June 2010. A total of National University of
110 clinical isolates were subjected to biofilm detection methods. Isolates were identified by standard Sciences and Technology
(NUST), Islamabad, Army
microbiological procedures. Biofilm detection was tested by TCP, TM and CRA. Antibiotic suscep- Medical College, Rawalpindi,
tibility test of biofilm producing bacteria was performed by using the Kirby-Bauer disc diffusion Pakistan
2
Professor of Pathology,
technique according to CLSI guidelines. Results: The TCP method was considered to be superior to Department of Microbiology,
TM and CRA. From the total of 110 clinical isolates, TCP method detected 22.7% as high, 41% mod- NUST, Islamabad, Army
Medical College, Rawalpindi,
erate and 36.3% as weak or non-biofilm producers. We have observed higher antibiotic resistance Pakistan
in biofilm producing bacteria than non-biofilm producers. Conclusion: We can conclude from our 3
FCPS Microbiology,
study that the TCP method is a more quantitative and reliable method for the detection of biofilm Department of Microbiology,
NUST, Islamabad, Army
forming microorganisms as compared to TM and CRA methods, and it can be recommended as a Medical College, Rawalpindi,
general screening method for detection of biofilm producing bacteria in laboratories. Pakistan
4
ST, Senior Technician,
Keywords: biofilms; bacteria; anti-bacterial agents. Department of Microbiology,
NUST, Islamabad, Army
Medical College, Rawalpindi,
Pakistan
INTRODUCTION a biofilm, as antibiotic resistance can increase
1,000 fold.3 According to a publication by the
Biofilms are defined as microbially derived ses-
National Institutes of Health, more than 80% of
sile communities characterized by the cells that
all infections involve biofilms.4 Biofilms are as-
are irreversibly attached to a substratum or to
sociated with many medical conditions includ-
each other. They are embedded in a matrix of
extracellular polymeric substances (EPS) they ing indwelling medical devices, dental plaque,
have produced, and exhibit an altered phe- upper respiratory tract infections, peritonitis,
notype with respect to growth rate and gene and urogenital infections.5 Both Gram-positive Submitted on: 01/03/2011
Approved on: 05/03/2011
transcription.1 Within a biofilm, bacteria com- and Gram-negative bacteria have the capability
municate with each other by production of to form biofilms. Bacteria commonly involved Correspondence to:
Afreenish Hassan
chemotactic particles or pheromones, a phe- include Enterococcus faecalis, Staphylococcus Department of Microbiology
nomenon called quorum sensing.2 Availability aureus, Staphylococcus epidermidis, Streptococ- National University of
Sciences and Technology,
of key nutrients, chemotaxis towards surface, cus viridans, Escherichia coli, Klebsiella pneu- Islamabad, Army Medical
motility of bacteria, surface adhesins and pres- moniae, Proteus mirabilis and Pseudomonas College, Rawalpindi,
Pakistan
ence of surfactants are some factors which in- aeruginosa.6 There are various methods to afreenish216a@yahoo.com
fluence biofilm formation.2 Microorganisms detect biofilm production. These include the
growing in a biofilm are intrinsically more re- Tissue Culture Plate (TCP),7 Tube method
We declare no conflict of
sistant to antimicrobial agents than planktonic (TM),8 Congo Red Agar method (CRA),9 bio- interest.
cells. High antimicrobial concentrations are luminescent assay,10 piezoelectric sensors,11
©2011 Elsevier Editora Ltda.
required to inactivate organisms growing in and fluorescent microscopic examination.12 All rights reserved.
305
BJID-4-JUN ARTE FINAL.indd 305 28/07/11 13:30
Evaluation of different detection methods of biofilm formation in the clinical isolates
We screened 110 organisms by three different methods, 0.2 mL of phosphate buffer saline (pH 7.2) four times. This
which could be used in a routine clinical laboratory, for removed free floating bacteria. Biofilm formed by bacteria
determining their ability to form biofilm. adherent to the wells were fixed by 2% sodium acetate and
stained by crystal violet (0.1%). Excess stain was removed by
OBJECTIVES using deionized water and plates were kept for drying. Opti-
cal density (OD) of stained adherent biofilm was obtained
The study was conducted to detect biofilm forming microor-
by using micro ELISA autoreader (model 680, Biorad, UK)
ganisms isolated from clinical specimens by three different
at wavelength 570 nm. The experiment was performed in
methods (TCP, TM, CRA) and to compare these methods
triplicate and repeated three times. The interpretation of
for biofilm detection.
biofilm production was done according to the criteria
MATERIALS AND METHODS of Stepanovic et al.14 (Table 2).
Tube method
Place and duration of the study Described by Christensen et al.,8 this is a qualitative
The study was conducted at the Department of Microbiol- method for biofilm detection. A loopful of test organ-
ogy, Army Medical College, National University of Sciences isms was inoculated in 10 mL of trypticase soy broth with
and Technology, Pakistan, from January 2010 to June 2010. 1% glucose in test tubes. The tubes were incubated at
37oC for 24 h. After incubation, tubes were decanted and
Selection of the isolates
washed with phosphate buffer saline (pH 7.3) and dried.
A total of 110 clinical isolates were subjected to biofilm de- Tubes were then stained with crystal violet (0.1%). Ex-
tection methods. Organisms were selected on the following cess stain was washed with deionized water. Tubes were
criteria: those isolated from the pus, intravenous and urinary
dried in inverted position. The scoring for tube method
catheter tips, urine, sputum and nasobronchial lavage speci-
was done according to the results of the control strains.
mens, and those showing increased resistance to commonly
Biofilm formation was considered positive when a visible
available antibiotics by Kirby-Bauer disc diffusion method.
film lined the wall and the bottom of the tube. The amount
Urinary catheter tips, intravenous catheter tips, nasobron-
of biofilm formed was scored as 1-weak/none, 2-moder-
chial lavage specimens and few of the pus specimens were
ate and 3-high/strong. The experiment was performed in
related to medical devices (Table 1). All of the specimens
triplicate and repeated three times.
were received from patients with nosocomial infections ad-
mitted to the hospital. Congo Red Agar method
Isolates were identified by standard microbiological pro-
Freeman et al.9 have described a simple qualitative method
cedures (Gram staining, colonial morphology, catalase test,
to detect biofilm production by using Congo Red Agar
cytochrome oxidase reaction, motility, biochemical tests).
(CRA) medium. CRA medium was prepared with brain
Reference strain of positive biofilm producer Staphylococcus
heart infusion broth (Oxoid, UK) 37 g/L, sucrose 50 g/L,
epidermidis ATCC 35984, Staphylococcus aureus ATCC
agar No. 1 (Oxoid, UK) 10 g/L and Congo Red indicator
35556, Pseudomonas aeruginosa ATCC 27853, Escherichia
(Oxoid, UK) 8 g/L. First Congo Red stain was prepared as
coli ATCC 35218 and Staphylococcus epidermidis ATCC
a concentrated aqueous solution and autoclaved (121oC for
12228 (non-slime producer) were used as control. Biofilm
15 minutes) separately from the other medium constitu-
detection was done by the following methods:
ents. Then it was added to the autoclaved brain heart infu-
Tissue culture plate method sion agar with sucrose at 55oC.5 CRA plates were inoculated
This quantitative test described by Christensen et al.7 is con- with test organisms and incubated at 37oC for 24 h aerobi-
sidered the gold-standard method for biofilm detection.13 cally. Black colonies with a dry crystalline consistency
Organisms isolated from fresh agar plates were inoculated in indicated biofilm production.5 The experiment was per-
10 mL of trypticase soy broth with 1% glucose. Broths were formed in triplicate and repeated three times.
incubated at 37oC for 24 h. The cultures were then diluted Antibiotic susceptibility test of biofilm produc-
1:100 with fresh medium. Individual wells of sterile 96 well- ing bacteria was done on Mueller Hinton agar (Oxoid,
flat bottom polystyrene tissue culture treated plates (Sigma- UK) using the following antibiotic discs: ampicillin,
Aldrich, Costar, USA) were filled with 200 µL of the diluted cotrimoxazole, ciprofloxacin, aztreonam, meropenem,
cultures. The control organisms were also incubated, diluted cefoperazone-sulbactam, chloramphenicol, vancomy-
and added to tissue culture plate. Negative control wells con- cin, erythromycin, amoxicillin-clavulanic acid, oxacil-
tained inoculated sterile broth. The plates were incubated at lin, linezolid, penicillin. All antibiotic discs were ob-
37oC for 24 h. After incubation, contents of each well were tained from Oxoid, UK. Escherichia coli ATCC 25922,
removed by gentle tapping. The wells were washed with Pseudomonas aeruginosa ATCC 27853 and Staphylococcus
306
BJID-4-JUN ARTE FINAL.indd 306 28/07/11 13:30
Hassan, Usman, Kaleem et al.
Table 1. Correlation of biofilm production of isolates and sensitivity test with the patient clinical condition and
type of medical device used
Organism Biofilm Pt. clinical Type of Sensitivity to
production condition/Type medical following
of infection device antibiotics
E. coli Strong Urine retention due Foley’s urinary Amikacin, meropenem,
to stroke catheter cefoperazone-sulbactam
S. epidermidis Strong Care of a metastatic Urinary catheter Vancomycin,
terminal disease linezolid, ciprofloxacin
E. coli Strong For relief of bladder Foley’s urinary catheter Aztreonam, meropenem
outlet obstruction due
to urethral stricture
E. faecalis Moderate Neurogenic bladder Foley’s urinary catheter Vancomycin, linezolid,
cotrimoxazole
E. coli Strong Benign prostatic Foley’s urinary catheter Ciprofloxacin,
hypertrophy aztreonam, meropenem
E. coli Strong Post-operation to Foley’s urinary catheter Aztreonam, meropenem,
monitor output ciprofloxacin
S. epidermidis Strong Urologic surgery Foley’s urinary catheter Vancomycin,
linezolid, ciprofloxacin
K. pneumoniae Strong Urine retention due Foley’s urinary catheter Cotrimoxazole,
to calculus disease ciprofloxacin,
ceftriaxone, meropenem
E. coli Strong Urine retention due to Foley’s urinary catheter Amikacin,
cerebrovascular disease ceftriaxone, meropenem
K. pneumoniae Strong Hip fracture surgery Foley’s urinary catheter Ciprofloxacin,
ceftriaxone, meropenem
E. coli Strong Post-operation Foley’s urinary catheter Meropenem, amikacin,
to monitor output aztreonam, ceftriaxone
S. epidermidis Strong Care for traumatic Foley’s urinary catheter Vancomycin, linezolid,
spinal cord injury cotrimoxazole,
ciprofloxacin
S. epidermidis Moderate Neurogenic bladder Foley’s urinary catheter Linezolid, ciprofloxacin,
cotrimoxazole
E. coli Strong Total abdominal Foley’s urinary catheter Ceftriaxone, meropenem
hysterectomy surgery
S. epidermidis Moderate Bladder outlet Foley’s urinary catheter Vancomycin, linezolid,
obstruction due cotrimoxazole,
to urethral stricture ciprofloxacin
E. coli Strong Urologic surgery Foley’s urinary catheter Meropenem, ceftriaxone
E. coli Strong Benign prostatic Foley’s urinary catheter Ciprofloxacin, aztreonam,
hypertrophy meropenem
K. pneumoniae Strong Palliative care in a Foley’s urinary catheter Aztreonam, meropenem
incontinent impaired
patient
S. epidermidis Strong Monitoring of central Central venous catheter Linezolid, ciprofloxacin
venous pressure
in an acutely ill patient
(Cont.)
Braz J Infect Dis 2011; 15(4):305-311 307
BJID-4-JUN ARTE FINAL.indd 307 28/07/11 13:30
Evaluation of different detection methods of biofilm formation in the clinical isolates
Table 1. Correlation of biofilm production of isolates and sensitivity test with the patient clinical condition and
type of medical device used
Organism Biofilm Pt. clinical Type of Sensitivity to
production condition/Type medical following
of infection device antibiotics
S. epidermidis Moderate Total parenteral nutrition Central venous catheter Vancomycin, linezolid,
cotrimoxazole,
ciprofloxacin
K. pneumoniae Strong Long term IV antibiotics Intravenous catheter Aztreonam,
cefoperazone-sulbactam,
meropenem
S. epidermidis Strong To correct electrolyte Intravenous catheter Vancomycin, linezolid,
imbalance, fluid Ciprofloxacin
replacement
S. aureus Moderate Blood transfusion Intravenous catheter Rifampicin, ciprofloxacin
vancomycin
S. epidermidis Strong Total parenteral Central venous catheter Linezolid,
nutrition cotrimoxazole
S. epidermidis Strong Monitoring of central Central venous catheter Vancomycin,
venous pressure in ciprofloxacin
a patient in ICU
K. pneumoniae Strong Pneumonia in a Endotracheal tube Meropenem, ceftriaxone,
patient on ventilator aztreonam
P. aeruginosa Strong Ventilator associated Endotracheal tube Cefoperazone-sulbactam,
pneumonia amikacin, meropenem
S. epidermidis Moderate Prosthetic joint infection Artificial joint Vancomycin, linezolid,
ciprofloxacin
S. epidermidis Strong Acute renal failure/ PD Catheter Vancomycin, linezolid,
peritoneal dialysis/ cotrimoxazole,
cellulites, peritonitis ciprofloxacin
S. aureus Moderate Prosthetic joint infection Artificial joint Rifampicin, linezolid,
vancomcin, erythromycin
Table 2. Interpretation of biofilm production RESULTS
Average OD value Biofilm production Among 110 isolates, TCP, the standard method, detected
25 as strong and 45 as moderate biofilm producers. The
≤ ODc / ODc < ~ ≤ 2x ODc Non/weak
majority of the organisms associated with biofilm produc-
2x ODc < ~ ≤ 4x ODc Moderate tion were S. epidermidis (37.1%) followed by E. coli (27.1%),
> 4x ODc Strong K. pneumoniae (15.7%), S. aureus (11.4%), E. faecalis
(4.2%) and P. aeruginosa (4.2%). Biofilm producing bac-
Optical density cut-off value (ODc) = average OD of negative teria were isolated from urine (30%) followed by urinary
control + 3x standard deviation (SD) of negative control. catheter tips (25.7%), pus (12.8%), sputum (11.4%), in-
travenous catheter tips (10%) and nasobronchial lavage
specimens (10%). Strong biofilm production was caused by
aureus ATCC 29213 were used as control strains. An- E. coli and S. epidermidis on Foley’s urinary catheter,
tibiotic susceptibility test was performed by using the mainly in immunocompromised patients, sensitive pre-
Kirby-Bauer disc diffusion technique according to CLSI dominantly to meropenem, aztreonam, vanomycin and
guidelines.15 linezolid. S. epidermidis was responsible for strong biofilm
308
BJID-4-JUN ARTE FINAL.indd 308 28/07/11 13:30
Hassan, Usman, Kaleem et al.
production in patients with intravenous catheters, sensi- We have observed higher antibiotic resistance in bio-
tive mostly to linezolid and vancomycin (Table 1). film producing bacteria than non-biofilm producers. By
By TM, the number of strong biofilm producers were the standard method (TCP), biofilm producing bacteria
21, moderate were 33 and weak or non-biofilm producers include the strong (25) and moderate biofilm producers
were 56. Very different results were observed by the CRA (45), and non-biofilm producing bacteria include non-
method, with which only four isolates showed black colo- biofilm producers (40) (Tables 4 and 5).
nies with crystalline appearance (Table 3).
Table 3. Screening of the isolates for biofilm formation by Tissue Culture Plate, Tube Method and Congo Red
Agar methods
Biofilm formation TCM n (%) TM n (%) CRA n (%)
High 25 (22.7) 21 (19) 4 (3.6)
No. of isolates (110)
Moderate 45 (41) 33 (30) 7 (6.3)
Weak/none 40 (36.3) 56 (51) 99 (90)
Table 4. Resistance pattern (%) of biofilm producing Gram-positive bacteria
Antimicrobial agent Biofilm producing Non-biofilm producing
Gram-positive organisms Gram-positive organisms
% %
Penicillin 100 100
Rifampicin 70 30
Ciprofloxacin 40 10
Erythromycin 40 20
Cotrimoxazole 30 25
Linezolid 0 0
Vancomycin 0 0
Table 5. Resistance pattern (%) of biofilm producing Gram-negative bacteria
Antimicrobial agent Biofilm producing Non-biofilm producing
Gram-negative organisms Gram-negative organisms
% %
Ampicillin 100 100
Ciprofloxacin 95 50
Cotrimoxazole 90 83
Aztreonam 90 50
Amikacin 64 37
Ceftriaxone 58 33
Cefoperazone-sulbactam 36 0
Meropenem 0 0
Braz J Infect Dis 2011; 15(4):305-311 309
BJID-4-JUN ARTE FINAL.indd 309 28/07/11 13:30
Evaluation of different detection methods of biofilm formation in the clinical isolates
Table 6. Diagnostic parameters of Tube method and Congo Red Agar method for biofilm detection
Screening Sensitivity Specificity Positive predictive Negative predictive Accuracy
method (%) (%) value (%) value (%) (%)
TM 73 92.5 94 66 80
CRA 11 92 73 37 41
Statistical analysis of tissue culture plate, tube and TM is 73% sensitive, 92.5% specific and 80% accurate for
Congo Red Agar methods biofilm detection. This method correlated well with TCP
The TCP method was considered the gold-standard for this for identifying strong biofilm producers, but it was hard to
study and compared with data from TM and CRA meth- differentiate between moderate, weak and non-biofilm pro-
ods. Parameters like sensitivity, specificity, negative predic- ducers due to the changeability in the results detected by
tive value, positive predictive value and accuracy were cal- different observers. In accordance with the preceding stud-
culated. True positives were biofilm producers by TCP, TM ies, TM cannot be suggested as general screening test to
and CRA method. False positive were biofilm producers by identify biofilm producing isolates.8,13
TM and CRA method and not by TCP method. False nega- In another study, Ruzicka et al.19 noted that out of 147 iso-
tive were the isolates which were non-biofilm producers by lates of S. epidermidis, TM detected biofilm formation in 79
(53.7%) and CRA detected in 64 (43.5%) isolates. They showed
TM and CRA but were producing biofilm by TCP method.
that TM is better for biofilm detection than CRA.19 Baqai
True negatives are those which were non biofilm producers
et al.20 tested TM to detect biofilm formation among uropatho-
by all the three methods. Sensitivity and specificity of TM
gens. According to their results, 75% of the isolates exhibited
was 73% and 92.5%, respectively. For CRA method, sensi-
biofilm formation.20 With the CRA method, 11 were found to
tivity and specificity remained low and were 11% and 92%,
be biofilm producing bacteria and 99 as non-biofilm produc-
respectively (Table 6).
ers. The CRA method showed very little correlation with the
other methods and parameters of sensitivity (11%), specificity
DISCUSSION
(92%) and accuracy (41%) were very low. By this method, three
Biofilm producing bacteria are responsible for many recal- isolates were found to be false positive and 62 were false nega-
citrant infections and are notoriously difficult to eradicate. tive. Knobloch et al.21 did not recommend the CRA method for
They exhibit resistance to antibiotics by various methods like biofilm detection in their study. Out of 128 isolates of S. aureus,
restricted penetration of antibiotic into biofilms, decreased CRA detected only 3.8% as biofilm producers as compared to
growth rate and expression of resistance genes.16 There are TCP which detected 57.1% as biofilm producing bacteria.21
various methods for biofilm detection.7-12 In this study we
evaluated 110 isolates by three screening methods for their CONCLUSION
ability to form biofilms.
We can conclude from our study that TCP is a quantitative
In our study, we have found that the majority of biofilm
and reliable method to detect biofilm forming microorgan-
producing bacteria was from urinary catheter tips (26.3%).
isms. When compared to TM and CRA methods, and TCP
Similarly, Donlan6 reported in his study the association of
can be recommended as a general screening method for de-
biofilm producing bacteria with urinary catheters. tection of biofilm producing bacteria in laboratories.
In the TCP method, the number of isolates showing bi-
ofilm formation was 70 (64.7%), and non or weak biofilm
producers were 40 (36.3%). Regional data from India also REFERENCES
showed that out of 152 isolates tested, the number of bio-
film producers identified by TCP method was 53.9 %, and 1. Donlan RM, Costerton W. Biofilms: Survival mechanisms of
clinically relevant Microorganisms. Clin Microbiol Rev 2002;
non-biofilm producers were 46%.13 We have performed the 15(2):167-93.
TCP method by addition of 1% glucose in trypticase soy 2. Thomas D, Day F. Biofilm formation by plant associated bacte-
broth. Addition of sugar helps in biofilm formation.16,17 ria. Ann Rev Microbiol 2007; 61:401-22
This was also reported by studies conducted by Mathur et 3. Stewart PS, Costerton JW. Antibiotic resistance of bacteria in
al.13 and Bose et al.18 biofilms. Lancet 2001; 358(9276):135-8.
4. Research on microbial biofilms (PA-03-047). NIH, National
Tube method detected 49% isolates as biofilm producers Heart, Lung, and Blood Institute. 2002-12-20.
and 51% as non-biofilm producers. By this method, three iso- 5. Reid G. Biofilms in infectious disease and on medical devices.
lates were found to be false positive and 19 were false negative. Int. J. Antimic Ag 1999; 11:223-6.
310
BJID-4-JUN ARTE FINAL.indd 310 28/07/11 13:30
Hassan, Usman, Kaleem et al.
6. Donlan RM. Biofilms and device-associated infections. Emerg 14. Stepanovic S, Vukovi D, Hola V et al. Quantification of biofilm
Infect Dis 2001; 7(2): 277-81. in microtiter plates: overview of testing conditions and practi-
7. Christensen GD, Simpson WA, Younger JA et al. Adherence cal recommendations for assessment of biofilm production by
of coagulase negative Staphylococci to plastic tissue cultures: Staphylococci. APMIS. 2007 115:891-9.
a quantitative model for the adherence of Staphylococci to 15. Bauer AW, Kirby M M, Sherris JC, Jurek M. Antibiotic sus-
medical devices. J Clin Microbiol 1995;22:996-1006. ceptibility testing by a standardized single method. Am J Clin
8. Christensen GD, Simpson WA, Bisno AL, Beachey EH. Adher- Pathol 1966; 45:493-6.
ence of slime producing strains of Staphylococcus epidermidis 16. Kim L. Riddle of biofilm resistance. Antimic Ag Chemother
to smooth surfaces. Infect Immun1982; 37:318-26. 2001; 45(4):999-1007.
9. Freeman J, Falkiner FR, Keane CT. New method for detecting 17. Eftikhar F, Speert DP. Biofilm formation by persistent and
slime production by coagulase negative staphylococci. J Clin non-persistent isolates of Staphylococcus epidermidis from a
Pathol 1989; 42:872-4. neonatal intensive care unit. J Hosp Infect 2009; 71(2):112-6.
10. Donlan RM, Murga R, Bell M et al. Protocol for detection of 18. Bose S, Khodke M, Basak S, Mallick SK. Detection of biofilm
biofilms on needleless connectors attached to central venous producing staphylococci: need of the hour. J Clin Diagn Res
catheters. J Clin Microbiol 2001; 39:750-3. 2009; 3:1915-20.
11. Aparna MS, Yadav S. Biofilms: microbes and disease. Braz 19. Ruzicka F, Hola V, Votava M et al. Biofilm detection and clini-
J Infect Dis 2008; 12(6):526-30. cal significance of Staphylococcus epidermidis isolates. Folia
12. Zufferey J, Rime B, Francioli P, Bille J. Simple method for rapid Microbiol (Praha) 2004; 49(5):596-600.
diagnosis of catheter associated infection by direct Acridine or- 20. Baqai R, Aziz M, Rasool G. Urinary tract infection in diabetic
ange staining of catheter tips. J Clin Microbiol 1988; 26:175-7. patients and biofilm formation of uropathogens. Infect Dis
13. Mathur T, Singhal S, Khan S, Upadhyay DJ, Fatma T, Rattan A. J Pakistan. 2008 17(1):7-9.
Detection of biofilm formation among the clinical isolates of 21. Knobloch JK, Horsetkotte MA, Rohde H, Mack D. Evaluation
staphylococci: an evaluation of three different screening meth- of different detection methods of biolfilm formation in Staphy-
ods. Indian J Med Microbiol 2006; 24(1) :25-9. lococcus aureus. Med Microbial Immunol 2002; 191(2):101-6.
Braz J Infect Dis 2011; 15(4):305-311 311
BJID-4-JUN ARTE FINAL.indd 311 28/07/11 13:30
View publication stats
You might also like
- 2022 Book ProtocolsForTheDiagnosisOfPigVDocument383 pages2022 Book ProtocolsForTheDiagnosisOfPigV谭硕100% (1)
- Textbook of Practical MicrobiologyDocument329 pagesTextbook of Practical MicrobiologyMara Maruca94% (50)
- Cytokine Bioassays PDFDocument383 pagesCytokine Bioassays PDFSubash Suba100% (1)
- 2 PMID 37992081 BiofilmDocument15 pages2 PMID 37992081 Biofilmajaya basnetNo ratings yet
- 2Document12 pages2Annu OjhaNo ratings yet
- Mycoplasma Pneumoniae: Prevalance in Navi Mumbai, India Biomedical. Indian Journal of Applied ResearchDocument4 pagesMycoplasma Pneumoniae: Prevalance in Navi Mumbai, India Biomedical. Indian Journal of Applied ResearchSpinalis 2017No ratings yet
- Tube Method and Congo Red Agar Versus Tissue CultuDocument7 pagesTube Method and Congo Red Agar Versus Tissue CultuMohanad TofahNo ratings yet
- Microbiological Laboratory Techniques - Clinical GateDocument18 pagesMicrobiological Laboratory Techniques - Clinical GateCamila Carla CapriglioniNo ratings yet
- Study of Extended Spectrum Beta Lactamases & Biofilm in Pseudomonas Species Isolated From Different Clinical SamplesDocument7 pagesStudy of Extended Spectrum Beta Lactamases & Biofilm in Pseudomonas Species Isolated From Different Clinical SamplesIJAR JOURNALNo ratings yet
- Prospective Multicentre Evaluation of The Direct NDocument5 pagesProspective Multicentre Evaluation of The Direct Npooja pandeyNo ratings yet
- BMS Iddt 2020 41Document8 pagesBMS Iddt 2020 41Guneyden GuneydenNo ratings yet
- 2022 Article 19844Document12 pages2022 Article 19844raphaNo ratings yet
- CV Abdul HafeezDocument7 pagesCV Abdul HafeezAbdul Hafeez KandhroNo ratings yet
- Diagnosing: Jericko MagistradoDocument25 pagesDiagnosing: Jericko MagistradoJericko MagistradoNo ratings yet
- Periodic Acid Schiff Reactions and General Tissue Morphology of Conventionally Processed Versus Two Rapid Microwave Processed TissuesDocument15 pagesPeriodic Acid Schiff Reactions and General Tissue Morphology of Conventionally Processed Versus Two Rapid Microwave Processed TissuesRestiyaniNo ratings yet
- HHS Public Access: Getting Started With Microbiome Analysis: Sample Acquisition To BioinformaticsDocument41 pagesHHS Public Access: Getting Started With Microbiome Analysis: Sample Acquisition To Bioinformatics@lsreshtyNo ratings yet
- Basic Microbiological TestsDocument13 pagesBasic Microbiological TestsyeabsraNo ratings yet
- Guidelinesfor Quick Applicationof Biochemical Teststo Identify Unknown BacteriaDocument19 pagesGuidelinesfor Quick Applicationof Biochemical Teststo Identify Unknown Bacteriara.rodriguezojedaNo ratings yet
- Microbiological Examination For Suspected BacteriaDocument8 pagesMicrobiological Examination For Suspected BacteriaAhmad Alfiansyah SaidNo ratings yet
- Journal of Biosafety and BiosecurityDocument5 pagesJournal of Biosafety and BiosecuritypratiwiNo ratings yet
- Antibio AbstractDocument2 pagesAntibio Abstractvaidyam78No ratings yet
- Laboratory Guide Methodologies For Antimicrobial Susceptibility TestingDocument19 pagesLaboratory Guide Methodologies For Antimicrobial Susceptibility TestingSharina Mae Quilaquiga FidelNo ratings yet
- Mag Hale MostafaDocument8 pagesMag Hale MostafaFranco SantinNo ratings yet
- First Priniciples of Clinical MicrobiologyDocument7 pagesFirst Priniciples of Clinical MicrobiologyDi MonizNo ratings yet
- 45 Supplement - 2 S99Document13 pages45 Supplement - 2 S99Mustaqeem DawarNo ratings yet
- Cells 10 00463 v2Document40 pagesCells 10 00463 v2Stephanie Nichole Ian CasemNo ratings yet
- Formulation and Evaluation of Acyclovir MicrocapsuDocument5 pagesFormulation and Evaluation of Acyclovir MicrocapsuRizQi FatmiyahNo ratings yet
- Principle and Techniques of Immunohistochemistry A ReviewDocument8 pagesPrinciple and Techniques of Immunohistochemistry A ReviewAyunda Novita RaniNo ratings yet
- Biocompatibility Test Methods - Pacific BioLabsDocument11 pagesBiocompatibility Test Methods - Pacific BioLabsAnil KumarNo ratings yet
- Coagulase Negative Staphylococcus Species : Resistance and Therapeutic Decisions at The Turn of The Novel MillenniumDocument6 pagesCoagulase Negative Staphylococcus Species : Resistance and Therapeutic Decisions at The Turn of The Novel MillenniumdrpklalNo ratings yet
- Maldi-Tof MsDocument12 pagesMaldi-Tof MsD. GandhirajNo ratings yet
- Association of Virulence Genes With Antibiotic Resistance in Pakistani Uropathogenic E. Coli IsolatesDocument6 pagesAssociation of Virulence Genes With Antibiotic Resistance in Pakistani Uropathogenic E. Coli Isolateskhurry testNo ratings yet
- A Practical Guide To ISO 10993-5 - Cytotoxicity - MDDI Medical Device and Diagnostic Industry News Products and SuppliersDocument4 pagesA Practical Guide To ISO 10993-5 - Cytotoxicity - MDDI Medical Device and Diagnostic Industry News Products and SuppliersVanessa DuzNo ratings yet
- FTIR Analysis of Natural and Synthetic Collagen: Applied Spectroscopy Reviews March 2018Document46 pagesFTIR Analysis of Natural and Synthetic Collagen: Applied Spectroscopy Reviews March 2018Imelda Irma IsauraNo ratings yet
- Jomb 2014 0043 PDFDocument9 pagesJomb 2014 0043 PDFGonzalez ArturoNo ratings yet
- B.sc. Medical Laboratory Sciences Medical Bacteriology (MLS 2413) (PDFDrive)Document101 pagesB.sc. Medical Laboratory Sciences Medical Bacteriology (MLS 2413) (PDFDrive)Isah MohammedNo ratings yet
- 12 Catheter TipDocument6 pages12 Catheter Tipselvakumar selvarajNo ratings yet
- 404-Article Text-2908-1-10-20230508Document14 pages404-Article Text-2908-1-10-20230508elbizco8No ratings yet
- An Assignment On Methods of Sterilization: Submitted ToDocument35 pagesAn Assignment On Methods of Sterilization: Submitted ToShahin Rahman 112700gmail.comNo ratings yet
- Detection of Antibiotic Resistance and Bio Film-Producing Ability of Staphylococcus Species in Clinical IsolatesDocument5 pagesDetection of Antibiotic Resistance and Bio Film-Producing Ability of Staphylococcus Species in Clinical Isolatesafnan jumaaNo ratings yet
- The Term Antibiotic Refers To Any Substance Produced by Any Microbial Entity That Is Detrimental To Other Other Microbial SpeciesDocument26 pagesThe Term Antibiotic Refers To Any Substance Produced by Any Microbial Entity That Is Detrimental To Other Other Microbial SpeciesPratibha PatilNo ratings yet
- Biosafety in Microbiological Laboratories M A LaysiasingaporeindiaDocument385 pagesBiosafety in Microbiological Laboratories M A Laysiasingaporeindiamohd farihan musthafa kamalNo ratings yet
- Anticancer Activities of Antimicrobial Bmkn2 Peptides Against Oral and Colon Cancer CellsDocument12 pagesAnticancer Activities of Antimicrobial Bmkn2 Peptides Against Oral and Colon Cancer CellselustosacamposNo ratings yet
- 1 s2.0 S2225411022000554 MainDocument11 pages1 s2.0 S2225411022000554 MaindanielwongtsNo ratings yet
- 30395-Article Text-56938-1-10-20220916Document8 pages30395-Article Text-56938-1-10-20220916Djim KARYOMNo ratings yet
- Evaluation of Diffusion and Dilution Methods To Determine The Antibacterial Activity of Plant ExtractsDocument7 pagesEvaluation of Diffusion and Dilution Methods To Determine The Antibacterial Activity of Plant ExtractsHusniya FaradisaNo ratings yet
- Wa0011.Document11 pagesWa0011.umayamajidNo ratings yet
- Integrating Molecular Biology Into The Veterinary CurriculumDocument17 pagesIntegrating Molecular Biology Into The Veterinary CurriculumSin PermahaNo ratings yet
- Dont KnowDocument5 pagesDont KnownawarajNo ratings yet
- PCR Optimization For Beginners A Step by Step GuideDocument23 pagesPCR Optimization For Beginners A Step by Step GuideĐặng Gia HoàngNo ratings yet
- Paso Corto 7 - Diagnostic Microbiology in Veterinary Dermatology-1Document10 pagesPaso Corto 7 - Diagnostic Microbiology in Veterinary Dermatology-1ABIGAIL AKEMI RODRIGUEZ CHAVEZNo ratings yet
- Guidance On Good Cell Culture PracticeDocument28 pagesGuidance On Good Cell Culture PracticeNyna KawlesNo ratings yet
- Diagnosis of Bacterial Infections (QH 29.02.2016)Document50 pagesDiagnosis of Bacterial Infections (QH 29.02.2016)rasool ghaffariNo ratings yet
- Screening and Selection of Industrially Important Microorganisms: A ReviewDocument7 pagesScreening and Selection of Industrially Important Microorganisms: A Reviewjeevalakshmanan29No ratings yet
- 1 s2.0 S1021949819300146 MainDocument11 pages1 s2.0 S1021949819300146 MainraraNo ratings yet
- BacterialisolationDocument10 pagesBacterialisolationMark SudieNo ratings yet
- Reviews:: Protein Cancer Biomarkers Serve MultiDocument11 pagesReviews:: Protein Cancer Biomarkers Serve MultiIoana CreangaNo ratings yet
- Comparison of Chemiluminescence Enzyme ImmunoassayDocument7 pagesComparison of Chemiluminescence Enzyme ImmunoassayDewi ArisandyNo ratings yet
- Diagnosis MikrobiologiDocument14 pagesDiagnosis Mikrobiologiarmelya aisyahNo ratings yet
- Analytical MicrobiologyFrom EverandAnalytical MicrobiologyFrederick KavanaghNo ratings yet
- WerwerwerDocument1 pageWerwerwerYanuar PrasetyoNo ratings yet
- DasdadasdDocument4 pagesDasdadasdYanuar PrasetyoNo ratings yet
- E.G. OP-11-Cherry - Pptx. For Your Presentation Code, Please Refer To The List of OralDocument2 pagesE.G. OP-11-Cherry - Pptx. For Your Presentation Code, Please Refer To The List of OralYanuar PrasetyoNo ratings yet
- Lai Et Al-2013-Arthritis & RheumatismDocument1 pageLai Et Al-2013-Arthritis & RheumatismYanuar PrasetyoNo ratings yet
- List of Oral Presenters: Time (WIB) Presentation Code Name RoomDocument2 pagesList of Oral Presenters: Time (WIB) Presentation Code Name RoomYanuar PrasetyoNo ratings yet
- Inhibitor-Based Methods For Detection of Plasmid-Mediated Ampc - Lactamases in Klebsiella SPP., Escherichia Coli, and Proteus MirabilisDocument5 pagesInhibitor-Based Methods For Detection of Plasmid-Mediated Ampc - Lactamases in Klebsiella SPP., Escherichia Coli, and Proteus MirabilisYanuar PrasetyoNo ratings yet
- Rounds: Case Study and Review of Autoimmune HepatitisDocument7 pagesRounds: Case Study and Review of Autoimmune HepatitisClaudia Naomi Ventura OrtizNo ratings yet
- Human Fertilization Is The Union of A Human Egg and SpermDocument7 pagesHuman Fertilization Is The Union of A Human Egg and Spermoxidalaj100% (1)
- Stem Cell Powerpoint With Teacher NotesDocument36 pagesStem Cell Powerpoint With Teacher NotesDELLA BLATAMANo ratings yet
- Influence of Antiretroviral Therapy On Maternal Eosinophil Levels During Pregnancy: A ReviewDocument14 pagesInfluence of Antiretroviral Therapy On Maternal Eosinophil Levels During Pregnancy: A ReviewKIU PUBLICATION AND EXTENSIONNo ratings yet
- Department of Internal Medicine: Non-Hodgkin LymphomaDocument26 pagesDepartment of Internal Medicine: Non-Hodgkin LymphomaHana FauziNo ratings yet
- Culture Media Culture Media PurposeDocument2 pagesCulture Media Culture Media Purposeangelique wongNo ratings yet
- The Complete Blood Cell Count A PowerfulDocument16 pagesThe Complete Blood Cell Count A PowerfulGabriele VitorNo ratings yet
- Repurposing Non Oncology Small Molecule Drugs To Improve - 2022 - Acta PharmaceDocument26 pagesRepurposing Non Oncology Small Molecule Drugs To Improve - 2022 - Acta PharmaceMohammed Shuaib AhmedNo ratings yet
- Rennets: MicrobialDocument35 pagesRennets: MicrobialJonathan CisternasNo ratings yet
- Fosbac Plus T (F+T) : in Broiler ProductionDocument25 pagesFosbac Plus T (F+T) : in Broiler Productionthanh ba matNo ratings yet
- The Histone Code HypothesisDocument5 pagesThe Histone Code HypothesisBrian D. StrahlNo ratings yet
- MicrosatellitesDocument24 pagesMicrosatellitesAhmed BioNo ratings yet
- Salmonella Pathogenicity Islands Encoding Type III Secretion SystemsDocument11 pagesSalmonella Pathogenicity Islands Encoding Type III Secretion SystemsIsabella GriffithNo ratings yet
- Introduction To Cell BiologyDocument43 pagesIntroduction To Cell BiologyEllemrac GageloniaNo ratings yet
- HAP Lab Quiz (Midterm)Document11 pagesHAP Lab Quiz (Midterm)shane.erica.penitNo ratings yet
- Evaluation of The Performance of Sysmex XN-3100 Automated Hematology Analyzer Regarding The Sysmex XE-2100 and Microscopic ExaminationDocument9 pagesEvaluation of The Performance of Sysmex XN-3100 Automated Hematology Analyzer Regarding The Sysmex XE-2100 and Microscopic ExaminationbalkisNo ratings yet
- Culture Media: For Industrial MicrobiologyDocument34 pagesCulture Media: For Industrial MicrobiologyKATHENo ratings yet
- What Are The HLA Genes?: Humans Have Three Main MHC Class I Genes, Known As HLA-A, HLA-B, and HLA-C. TheDocument2 pagesWhat Are The HLA Genes?: Humans Have Three Main MHC Class I Genes, Known As HLA-A, HLA-B, and HLA-C. ThepoojaNo ratings yet
- Monoclonal AntibodiesDocument35 pagesMonoclonal Antibodiespreetylyall100% (5)
- Mark Blood BankDocument20 pagesMark Blood BankMark Lester MoralesNo ratings yet
- Review Sheet On 7.2Document9 pagesReview Sheet On 7.2KJannat Mahmood owaitanNo ratings yet
- QUALITY MANUAL v008 FinalDocument22 pagesQUALITY MANUAL v008 FinalPERICODELOSPALOTESNo ratings yet
- Ingles Area1 Jan2016Document4 pagesIngles Area1 Jan2016Eugênio RegoNo ratings yet
- Litareture ReviewDocument3 pagesLitareture ReviewShowkatul IslamNo ratings yet
- Dr.R.a.reizkhi TugasDocument13 pagesDr.R.a.reizkhi TugasReizkhi FitriyanaNo ratings yet
- Principles of ImmunizationDocument4 pagesPrinciples of ImmunizationDoc Prince CaballeroNo ratings yet
- WES WGS Brochure Pages V2.1eng Web 20201019Document16 pagesWES WGS Brochure Pages V2.1eng Web 20201019drumerNo ratings yet
- Booster Mock Test For NEETDocument15 pagesBooster Mock Test For NEETDrNo ratings yet
- Virology Respiratory VirusesDocument6 pagesVirology Respiratory VirusesHisham ChomanyNo ratings yet