Professional Documents
Culture Documents
Kimball Et Al 2018, Figure 9
Kimball Et Al 2018, Figure 9
Uploaded by
Abby KimballCopyright:
Available Formats
You might also like
- 1 - Light Plot - Overhead - Hairspray - B03Document1 page1 - Light Plot - Overhead - Hairspray - B03Mike WoodNo ratings yet
- Gigabyte GA-Z77-DS3H Rev 1.0 BoardView PDFDocument1 pageGigabyte GA-Z77-DS3H Rev 1.0 BoardView PDFKhánh GiaNo ratings yet
- Column Shop DrawingDocument1 pageColumn Shop DrawingdantevariasNo ratings yet
- Exaudiat Te DominusDocument5 pagesExaudiat Te DominusCatalina ʚïɞNo ratings yet
- PRC Ple Review 021318Document301 pagesPRC Ple Review 021318Maria Cecilia Luz Romero67% (6)
- Carpark Layout PlanDocument1 pageCarpark Layout PlanP CyNo ratings yet
- GA H77 DS3H R1.0 ăÎÍDocument1 pageGA H77 DS3H R1.0 ăÎÍMinh Nguyen PhuNo ratings yet
- Dragon Ball Z - Cha-La Head-Cha-LaDocument3 pagesDragon Ball Z - Cha-La Head-Cha-LaJI RANo ratings yet
- 740 - Opsb3200-3215-3218Document3 pages740 - Opsb3200-3215-3218Jessica JacobNo ratings yet
- v3m BlockedDocument1 pagev3m BlockedRaúl OjedaNo ratings yet
- 8013 Tsa 016 DW 1452 C 108Document1 page8013 Tsa 016 DW 1452 C 108Marty TuberNo ratings yet
- ReleaseNote FileList of X415DA 2009 X64 V1.00Document7 pagesReleaseNote FileList of X415DA 2009 X64 V1.00Karen AdanaqueNo ratings yet
- 033 - de Colores - C - 02 PDFDocument1 page033 - de Colores - C - 02 PDFJan WillemsNo ratings yet
- PVI 6000 TL OUTD W Quick Installation Guide en RevADocument2 pagesPVI 6000 TL OUTD W Quick Installation Guide en RevA王伯恩No ratings yet
- A-103 Ground Floor Plan Revised1346586168420Document1 pageA-103 Ground Floor Plan Revised1346586168420zubair khanNo ratings yet
- Ta41 40007eDocument2 pagesTa41 40007eJuan CoronelNo ratings yet
- 51 BEL113 - Longstreet Dixie - Abel - Page - 0033 F Horn 2Document1 page51 BEL113 - Longstreet Dixie - Abel - Page - 0033 F Horn 2Giuseppe RufiniNo ratings yet
- A-07-003a - ENLARGED FIRST FLOOR PLAN (TYPICAL PER BUILDING)Document1 pageA-07-003a - ENLARGED FIRST FLOOR PLAN (TYPICAL PER BUILDING)zizojundiNo ratings yet
- L01 MBLC D05 Fad DWG 07PZ100 ZZ Ele1104 R00Document1 pageL01 MBLC D05 Fad DWG 07PZ100 ZZ Ele1104 R00Asif SafiNo ratings yet
- Pmi 635Document3 pagesPmi 635api-3703813No ratings yet
- El Derecho de Vivir en Paz - Tenor 1Document3 pagesEl Derecho de Vivir en Paz - Tenor 1santiago cerdaNo ratings yet
- SPECIALZ Animenz - Original Artist King GnuDocument6 pagesSPECIALZ Animenz - Original Artist King GnujoemwirichiaNo ratings yet
- Passacaglia - Handel 4 RecordersDocument3 pagesPassacaglia - Handel 4 RecordersRiccardo PopNo ratings yet
- Buseko WarehouseDocument1 pageBuseko Warehouselenganji simwandaNo ratings yet
- SMS Nobreak Sen T03395-02 Manager 1400 USBDocument1 pageSMS Nobreak Sen T03395-02 Manager 1400 USBSamuel LopesNo ratings yet
- Basisanschlussplan MS4 Sport HPI DDU7 Software Release 36Document1 pageBasisanschlussplan MS4 Sport HPI DDU7 Software Release 36Sebastian BryceNo ratings yet
- Footlose: Kenny LogginsDocument2 pagesFootlose: Kenny LogginsMarco PepeNo ratings yet
- Inverex 3525, LM 358 Circuit DaigramDocument1 pageInverex 3525, LM 358 Circuit DaigramAmjad HussainNo ratings yet
- Petronas Carigali SDN BHD: Document Review StatusDocument1 pagePetronas Carigali SDN BHD: Document Review StatusMohd KhaidirNo ratings yet
- 87ae5500 Cyr - Uty%Document1 page87ae5500 Cyr - Uty%Narendra GadeNo ratings yet
- SRI JAYAJOTHI UNIT OE STRING LAYOUT 1-ModelDocument1 pageSRI JAYAJOTHI UNIT OE STRING LAYOUT 1-ModelDIVAKARNo ratings yet
- Esquema Elétrico Pci Principal: From TransDocument8 pagesEsquema Elétrico Pci Principal: From TransLuiz Clemente PimentaNo ratings yet
- C-Hmi Server Client: ABB ABBDocument1 pageC-Hmi Server Client: ABB ABBmahmoud sheblNo ratings yet
- Fsa 1210 F2Document1 pageFsa 1210 F2Nagamani ManiNo ratings yet
- To taufiqDocument1 pageTo taufiqJevarajan GanasanNo ratings yet
- MIC-EST-PE-08-R01-RAMPA (1)Document1 pageMIC-EST-PE-08-R01-RAMPA (1)Lucas MarçalNo ratings yet
- Il Vento D'oro Big Band Arrangement-PianoDocument9 pagesIl Vento D'oro Big Band Arrangement-PianoRobin SveginNo ratings yet
- 07-22-3971 - USB SchematicDocument1 page07-22-3971 - USB SchematicHamiltonNo ratings yet
- MoliendoDocument1 pageMoliendofioNo ratings yet
- Cooling Fand Corolla 2010Document1 pageCooling Fand Corolla 2010Alfredo jose Medina revattaNo ratings yet
- Bidp SDH 2022 Ar 119 DtcotDocument1 pageBidp SDH 2022 Ar 119 DtcotJeffNo ratings yet
- DWG PDF 20.04.2022Document1 pageDWG PDF 20.04.2022Sanjan SameerNo ratings yet
- Ews DrainageDocument1 pageEws Drainagetahirshaikh143No ratings yet
- F F+ B GM C: Som: Ritmo: TempoDocument1 pageF F+ B GM C: Som: Ritmo: TempoAlex OliveiraNo ratings yet
- Screenshot 2022-10-20 at 3.59.19 PMDocument4 pagesScreenshot 2022-10-20 at 3.59.19 PMphearath.moonengineeringNo ratings yet
- (Superpartituras - Com.br) EvidenciasDocument1 page(Superpartituras - Com.br) EvidenciasCesar SilvaNo ratings yet
- (Superpartituras - Com.br) EvidenciasDocument1 page(Superpartituras - Com.br) EvidenciasPedro Henrique Pezzi BonattoNo ratings yet
- (Superpartituras - Com.br) EvidenciasDocument1 page(Superpartituras - Com.br) EvidenciasEliab Miranda AbNo ratings yet
- F F+ B GM C: Som: Ritmo: TempoDocument1 pageF F+ B GM C: Som: Ritmo: TempoBenicio MoriNo ratings yet
- Chitaozinho e Xororo - EvidenciasDocument1 pageChitaozinho e Xororo - EvidenciasBeatriz CarvalhoNo ratings yet
- (Superpartituras - Com.br) EvidenciasDocument1 page(Superpartituras - Com.br) EvidenciasMarco A. P. AmmirabileNo ratings yet
- Chitaozinho e Xororo Evidencias PDFDocument1 pageChitaozinho e Xororo Evidencias PDFVinicius MartinsNo ratings yet
- Chitaozinho e Xororo Evidencias PDFDocument1 pageChitaozinho e Xororo Evidencias PDFPHERNANDO E KAIKENo ratings yet
- La Chocolata Azinhaga-Saxofone AltoDocument1 pageLa Chocolata Azinhaga-Saxofone Altoinesdc5No ratings yet
- Legend Barangay Boundary Barangay Road: A C A B C D E FDocument1 pageLegend Barangay Boundary Barangay Road: A C A B C D E FJoel Rodello B DugeniaNo ratings yet
- Dspa595 SchDiagram OperationDocument1 pageDspa595 SchDiagram OperationAndrew Chan YanNo ratings yet
- Est-Casa Club Aires Del Batan-E2Document1 pageEst-Casa Club Aires Del Batan-E2ISMAEL MONTIELNo ratings yet
- Bidp SDH 2022 Ar 117 DTCMCDocument1 pageBidp SDH 2022 Ar 117 DTCMCJeffNo ratings yet
- Suite Violino IDocument2 pagesSuite Violino IDaniel PivaNo ratings yet
- Oh DarlingDocument3 pagesOh DarlingJoão PedroNo ratings yet
- Kimball Et Al 2019, Figure 7Document1 pageKimball Et Al 2019, Figure 7Abby KimballNo ratings yet
- Oko Et Al 2019, Figure 8Document1 pageOko Et Al 2019, Figure 8Abby KimballNo ratings yet
- Kimball Et Al 2019, Figure 2Document1 pageKimball Et Al 2019, Figure 2Abby KimballNo ratings yet
- Kimball Et Al 2019, Figure 3Document1 pageKimball Et Al 2019, Figure 3Abby KimballNo ratings yet
- Neuwelt Et Al, Figure 2Document1 pageNeuwelt Et Al, Figure 2Abby KimballNo ratings yet
- Kimball Et Al 2019, Figure 4Document1 pageKimball Et Al 2019, Figure 4Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 5Document1 pageKimball Et Al 2018, Figure 5Abby KimballNo ratings yet
- Kimball Et Al 2019, Figure 1Document1 pageKimball Et Al 2019, Figure 1Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 8Document1 pageKimball Et Al 2018, Figure 8Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 7Document1 pageKimball Et Al 2018, Figure 7Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 1Document1 pageKimball Et Al 2018, Figure 1Abby KimballNo ratings yet
- Bullock Et Al 2018, Supplemental Figure 5Document1 pageBullock Et Al 2018, Supplemental Figure 5Abby KimballNo ratings yet
- Graph Features:: Events Sampled Per File: 9,141 Minimum Cluster Size: 5% Group Assignment: CorrectDocument1 pageGraph Features:: Events Sampled Per File: 9,141 Minimum Cluster Size: 5% Group Assignment: CorrectAbby KimballNo ratings yet
- Berger Et Al 2019, Figure 2Document1 pageBerger Et Al 2019, Figure 2Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 2Document1 pageKimball Et Al 2018, Figure 2Abby KimballNo ratings yet
- Bullock Et Al 2018, Figure 6Document1 pageBullock Et Al 2018, Figure 6Abby KimballNo ratings yet
- Hospital Pharmacy Cover LetterDocument7 pagesHospital Pharmacy Cover Letterykzdmfajd100% (2)
- Borax PDFDocument4 pagesBorax PDFZi LaNo ratings yet
- Lobes of The Brain - FrontalDocument12 pagesLobes of The Brain - FrontalLoccos SienaNo ratings yet
- Make Every Session Count PDFDocument209 pagesMake Every Session Count PDFEgylany LanyNo ratings yet
- Xylocaine Dentaland AdrenalineDocument7 pagesXylocaine Dentaland Adrenalinefawwaz FofoNo ratings yet
- CP On AnginaDocument21 pagesCP On AnginaNavpreet KaurNo ratings yet
- A Classification Tree To Assist With Routine Scoring of The Clinical Frailty ScaleDocument6 pagesA Classification Tree To Assist With Routine Scoring of The Clinical Frailty ScaleOlga GuerreroNo ratings yet
- Physical Examination by DRDocument25 pagesPhysical Examination by DRapi-3739910100% (2)
- Seht Card Hospital ListDocument33 pagesSeht Card Hospital Listlucky xenNo ratings yet
- Analytical Method Technology Transfer Is Pe GuideDocument46 pagesAnalytical Method Technology Transfer Is Pe Guidevenkat_du2000100% (1)
- Fisher 2009Document6 pagesFisher 2009Maryanne CunhaNo ratings yet
- Paul Ramesh Forensic Neuro Psychological InterviewDocument43 pagesPaul Ramesh Forensic Neuro Psychological Interviewnonam2100% (2)
- Pacemaker PamphletDocument2 pagesPacemaker Pamphletapi-355079078No ratings yet
- FSC Extra ENT and Tuning Fork January 11-Th 2018 Internet ConferenceDocument21 pagesFSC Extra ENT and Tuning Fork January 11-Th 2018 Internet Conferencemarc800No ratings yet
- Supply Chain Management in HospitalsDocument6 pagesSupply Chain Management in HospitalsAbhra100% (10)
- BNF - 76 - British - National - Formulary - Septem-357-358 PhenobarbtalDocument2 pagesBNF - 76 - British - National - Formulary - Septem-357-358 Phenobarbtalnur aisahNo ratings yet
- Prostate, Lung, Colorectal and Ovarian Cancer Screening Trial Medication Use QuestionnaireDocument3 pagesProstate, Lung, Colorectal and Ovarian Cancer Screening Trial Medication Use QuestionnaireGeede AlfonsoNo ratings yet
- Notes To Remember: For Clinical Ophthalmology MCQ ExamsDocument89 pagesNotes To Remember: For Clinical Ophthalmology MCQ ExamsMohamed Gaber100% (1)
- Bronchopulmonary Dysplasia: An Update: Anita Bhandari and Vineet BhandariDocument5 pagesBronchopulmonary Dysplasia: An Update: Anita Bhandari and Vineet BhandariPatricius Berry Harsono PutraNo ratings yet
- Cumitech 23 - Infections of The Skin and Subcutaneous Tissues PDFDocument16 pagesCumitech 23 - Infections of The Skin and Subcutaneous Tissues PDFAnjali MohanNo ratings yet
- Sucralfate PDFDocument2 pagesSucralfate PDFJoshua Christian Penggele100% (1)
- Prioritization of Health ProblemsDocument2 pagesPrioritization of Health ProblemsrorenNo ratings yet
- SL 1 MPT 23Document2 pagesSL 1 MPT 23Samidul IslamNo ratings yet
- RSI For Nurses ICUDocument107 pagesRSI For Nurses ICUAshraf HusseinNo ratings yet
- CV - DR Andrew Balyeku - June 2018 BDocument4 pagesCV - DR Andrew Balyeku - June 2018 BIga_DanNo ratings yet
- Ascorbic Acid InjectionDocument6 pagesAscorbic Acid InjectionMaria NorilynNo ratings yet
- Students With Learning Disabilities and InclusionDocument5 pagesStudents With Learning Disabilities and InclusionEditor IJTSRD100% (3)
- Asbestoswise ASEA Information SheetsDocument32 pagesAsbestoswise ASEA Information SheetsHeung Yeung CheukNo ratings yet
- Black Seed Nigella Sativa A Cure To Every DiseaseDocument6 pagesBlack Seed Nigella Sativa A Cure To Every DiseaseShahmeer KhanNo ratings yet
Kimball Et Al 2018, Figure 9
Kimball Et Al 2018, Figure 9
Uploaded by
Abby KimballOriginal Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Kimball Et Al 2018, Figure 9
Kimball Et Al 2018, Figure 9
Uploaded by
Abby KimballCopyright:
Available Formats
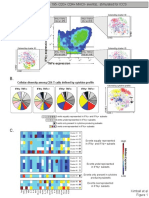
A.
CD11b CD11c Gr-1 PD-L1 Tbet
Gated on: CD4+
Some markers identified visually by
B6 #1
T cells in FlowJo
viSNE are not significantly different.
viSNE
**
IL10KO #1
B.
** 150
Median Intensity
***
ns
*
100
viSNE plots Altered location of CD4+ T cells, according 50
display only to tSNE1 and tSNE2, suggest B6
CD4+ T cell islands IL10KO 0
changes in phenotype of CD4+ T cells.
CD11b CD11c Gr-1 PD-L1 Tbet
C. B6
D. Focused analysis on CD4+ T cell clusters
CD11c CTLA4
Lag3
PhenoGraph defined six
subtypes of CD4+ T cells. PhenoGraph-defined
PhenoGraph
clusters differ in expression
Naive CD4+ T cells, (cluster #16) across multiple parameters.
% CD4+ T cells =16.3% Regulatory CD4+ T cells, (cluster #7)
CD11b ICOS IRF4
Effector CD4+ T cells A, (cluster #12)
IL10KO Effector CD4+ T cells B, (cluster #8)
Effector CD4+ T cells C, (cluster #28)
Effector CD4+ T cells D, (cluster #18) Certain cellular markers
are exclusively positive in
Effector CD4+ T cells A, (cluster #12)
Average of all events
IL10KO mice show IL10KO dominant clusters
Effector CD4+ T cells B, (cluster #8)
increased CD4+ T cells Effector CD4+ T cells C, (cluster #28) (cluster #8 and #28).
and altered distribution
Cluster #524
Individual Events
B6 biased
of CD4+ T cell subtypes. CD4+ T cell clusters that differ in
% CD4+ T cells =30.2%
abundance between conditions.
E. F. Line graphs of a cluster’s median
expression can be used to identify
cell type and phenotype subset.
Cluster #520
Individual Events
8 Cluster #515 Ki6
7 CD1
1b
MH
-1
Median Expression
Sca CI
S
PD1
6
The inner ring indicates core
ICO
IL10KO biased
X-shift
CD3
IRF4
4 CD4+ T cells
markers expressed on all
Tim3
Lag3
#515
1 L
2
44
PD
CD4+ T cells.
CD
Tb CD4 TR
Average of all events e
H.
t GI
Tbet
Tim3
Lag3
IkBa
CD19
NKp46
Ki67
PDL2
CD64
PDL1
IRF4
Sig F
FoxP3
KLRG1
Bcatenin
CD127
CD11c
GITR
MHCI
CD69
CD8
CD11b
CD25
Gr-1
p4eBP1
CD3
CTLA4
PD1
Sca1
CD45
CD44
CD4
CD117
MHCII
ICOS
CTLA4
100 Parameters
8 Cluster #520 IRF
4
-1 MH
Sca
CD
CI
Cellular expression
Median Expression
6
The outer ring defines
MHCII
Gr-1
across all parameters 4
Dendritic cells
X-shift can define the average #520
accessory phenotypes,
showing data from
CD
2
44
11
cellular expression across all
CD
with characteristics
Ki
CD11b
a
c
IkB
67
ten individual cells
parameters (here the most unique to this cell cluster,
Tbet
Tim3
Lag3
IkBa
CD19
NKp46
Ki67
PDL2
CD64
PDL1
IRF4
Sig F
FoxP3
KLRG1
Bcatenin
CD127
CD11c
GITR
MHCI
CD69
CD8
CD11b
CD25
Gr-1
p4eBP1
CD3
CTLA4
PD1
Sca1
CD45
CD44
CD4
CD117
MHCII
ICOS
(in cluster #520) Parameters
IL10KO biased cluster). not consistently present
highlights conserved,
among all cell subsets.
variable phenotypes.
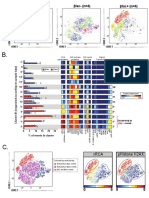
G. CD11b CD11c Gr-1 PD-L1 Tbet H. Node #1, CD4+ T cells
#26
5 ** ns
B6 #1
#1
ns
**
Median Intensity
Changed color on a node 4 ns
indicates altered expression
SPADE
#26 3
IL10KO #1
#1 between conditions.
2
B6
1
1.0 6.5 0.5 6.8 0.3 4.5 0.8 5.8 0.0 4.3
Median expression values IL10KO
for a single node can be 0
Selected CD4+ T cell nodes assessed for significance. CD11b CD11c Gr-1 PDL1 Tbet
K.
Number of model features #82228 IRF4 #82254 IRF4 #82228, PDL1 #82267, CD11b
I. J.
82233 82253
0 2 2 4 4 5 14 26 45 93 206 382 634
**** **** **** ****
Median Expression
Median Expression
Median Expression
Median Expression
82256
3 3 4 6
100
Cross Validation Error Rate Citrus Settings: 82248
3
82239
Feature False Discovery Rate Events sampled per file: 9,141
82261
2 2 4
cv.min
cv.1se
Minimum cluster size: 5%
Group assignment: Correct IRF4 2
PDL1
82265
cv.fdr.constrained
Cluster characterization: Medians 1 1 2
1
82264
Model Cross Validation Error Rate
82228 82249
cv.min model has 10% CD11b
75
82254 82250
0
82258
0
Citrus
0
82267
0
cross validation error rate
6
O
IL 6
O
IL 6
O
IL 6
O
82268 82222
B
B
B
K
82263
K
82260
10
10
10
10
82259
IL
82266
50
82237
82251
82262
82257
#82228
#82254
#82228
#82267
82252
82247
82255
25
82245
82241
IRF4 IRF4 PDL1 CD11b
82217
82220
Population: Background Cluster
Citrus identified 4 features that predict
0
Figure 9
7.15 6.36 5.56 4.77 3.97 3.18 2.38 1.59 0.79 0.00
Regularization Threshold
differences between experimental groups. Kimball et al
You might also like
- 1 - Light Plot - Overhead - Hairspray - B03Document1 page1 - Light Plot - Overhead - Hairspray - B03Mike WoodNo ratings yet
- Gigabyte GA-Z77-DS3H Rev 1.0 BoardView PDFDocument1 pageGigabyte GA-Z77-DS3H Rev 1.0 BoardView PDFKhánh GiaNo ratings yet
- Column Shop DrawingDocument1 pageColumn Shop DrawingdantevariasNo ratings yet
- Exaudiat Te DominusDocument5 pagesExaudiat Te DominusCatalina ʚïɞNo ratings yet
- PRC Ple Review 021318Document301 pagesPRC Ple Review 021318Maria Cecilia Luz Romero67% (6)
- Carpark Layout PlanDocument1 pageCarpark Layout PlanP CyNo ratings yet
- GA H77 DS3H R1.0 ăÎÍDocument1 pageGA H77 DS3H R1.0 ăÎÍMinh Nguyen PhuNo ratings yet
- Dragon Ball Z - Cha-La Head-Cha-LaDocument3 pagesDragon Ball Z - Cha-La Head-Cha-LaJI RANo ratings yet
- 740 - Opsb3200-3215-3218Document3 pages740 - Opsb3200-3215-3218Jessica JacobNo ratings yet
- v3m BlockedDocument1 pagev3m BlockedRaúl OjedaNo ratings yet
- 8013 Tsa 016 DW 1452 C 108Document1 page8013 Tsa 016 DW 1452 C 108Marty TuberNo ratings yet
- ReleaseNote FileList of X415DA 2009 X64 V1.00Document7 pagesReleaseNote FileList of X415DA 2009 X64 V1.00Karen AdanaqueNo ratings yet
- 033 - de Colores - C - 02 PDFDocument1 page033 - de Colores - C - 02 PDFJan WillemsNo ratings yet
- PVI 6000 TL OUTD W Quick Installation Guide en RevADocument2 pagesPVI 6000 TL OUTD W Quick Installation Guide en RevA王伯恩No ratings yet
- A-103 Ground Floor Plan Revised1346586168420Document1 pageA-103 Ground Floor Plan Revised1346586168420zubair khanNo ratings yet
- Ta41 40007eDocument2 pagesTa41 40007eJuan CoronelNo ratings yet
- 51 BEL113 - Longstreet Dixie - Abel - Page - 0033 F Horn 2Document1 page51 BEL113 - Longstreet Dixie - Abel - Page - 0033 F Horn 2Giuseppe RufiniNo ratings yet
- A-07-003a - ENLARGED FIRST FLOOR PLAN (TYPICAL PER BUILDING)Document1 pageA-07-003a - ENLARGED FIRST FLOOR PLAN (TYPICAL PER BUILDING)zizojundiNo ratings yet
- L01 MBLC D05 Fad DWG 07PZ100 ZZ Ele1104 R00Document1 pageL01 MBLC D05 Fad DWG 07PZ100 ZZ Ele1104 R00Asif SafiNo ratings yet
- Pmi 635Document3 pagesPmi 635api-3703813No ratings yet
- El Derecho de Vivir en Paz - Tenor 1Document3 pagesEl Derecho de Vivir en Paz - Tenor 1santiago cerdaNo ratings yet
- SPECIALZ Animenz - Original Artist King GnuDocument6 pagesSPECIALZ Animenz - Original Artist King GnujoemwirichiaNo ratings yet
- Passacaglia - Handel 4 RecordersDocument3 pagesPassacaglia - Handel 4 RecordersRiccardo PopNo ratings yet
- Buseko WarehouseDocument1 pageBuseko Warehouselenganji simwandaNo ratings yet
- SMS Nobreak Sen T03395-02 Manager 1400 USBDocument1 pageSMS Nobreak Sen T03395-02 Manager 1400 USBSamuel LopesNo ratings yet
- Basisanschlussplan MS4 Sport HPI DDU7 Software Release 36Document1 pageBasisanschlussplan MS4 Sport HPI DDU7 Software Release 36Sebastian BryceNo ratings yet
- Footlose: Kenny LogginsDocument2 pagesFootlose: Kenny LogginsMarco PepeNo ratings yet
- Inverex 3525, LM 358 Circuit DaigramDocument1 pageInverex 3525, LM 358 Circuit DaigramAmjad HussainNo ratings yet
- Petronas Carigali SDN BHD: Document Review StatusDocument1 pagePetronas Carigali SDN BHD: Document Review StatusMohd KhaidirNo ratings yet
- 87ae5500 Cyr - Uty%Document1 page87ae5500 Cyr - Uty%Narendra GadeNo ratings yet
- SRI JAYAJOTHI UNIT OE STRING LAYOUT 1-ModelDocument1 pageSRI JAYAJOTHI UNIT OE STRING LAYOUT 1-ModelDIVAKARNo ratings yet
- Esquema Elétrico Pci Principal: From TransDocument8 pagesEsquema Elétrico Pci Principal: From TransLuiz Clemente PimentaNo ratings yet
- C-Hmi Server Client: ABB ABBDocument1 pageC-Hmi Server Client: ABB ABBmahmoud sheblNo ratings yet
- Fsa 1210 F2Document1 pageFsa 1210 F2Nagamani ManiNo ratings yet
- To taufiqDocument1 pageTo taufiqJevarajan GanasanNo ratings yet
- MIC-EST-PE-08-R01-RAMPA (1)Document1 pageMIC-EST-PE-08-R01-RAMPA (1)Lucas MarçalNo ratings yet
- Il Vento D'oro Big Band Arrangement-PianoDocument9 pagesIl Vento D'oro Big Band Arrangement-PianoRobin SveginNo ratings yet
- 07-22-3971 - USB SchematicDocument1 page07-22-3971 - USB SchematicHamiltonNo ratings yet
- MoliendoDocument1 pageMoliendofioNo ratings yet
- Cooling Fand Corolla 2010Document1 pageCooling Fand Corolla 2010Alfredo jose Medina revattaNo ratings yet
- Bidp SDH 2022 Ar 119 DtcotDocument1 pageBidp SDH 2022 Ar 119 DtcotJeffNo ratings yet
- DWG PDF 20.04.2022Document1 pageDWG PDF 20.04.2022Sanjan SameerNo ratings yet
- Ews DrainageDocument1 pageEws Drainagetahirshaikh143No ratings yet
- F F+ B GM C: Som: Ritmo: TempoDocument1 pageF F+ B GM C: Som: Ritmo: TempoAlex OliveiraNo ratings yet
- Screenshot 2022-10-20 at 3.59.19 PMDocument4 pagesScreenshot 2022-10-20 at 3.59.19 PMphearath.moonengineeringNo ratings yet
- (Superpartituras - Com.br) EvidenciasDocument1 page(Superpartituras - Com.br) EvidenciasCesar SilvaNo ratings yet
- (Superpartituras - Com.br) EvidenciasDocument1 page(Superpartituras - Com.br) EvidenciasPedro Henrique Pezzi BonattoNo ratings yet
- (Superpartituras - Com.br) EvidenciasDocument1 page(Superpartituras - Com.br) EvidenciasEliab Miranda AbNo ratings yet
- F F+ B GM C: Som: Ritmo: TempoDocument1 pageF F+ B GM C: Som: Ritmo: TempoBenicio MoriNo ratings yet
- Chitaozinho e Xororo - EvidenciasDocument1 pageChitaozinho e Xororo - EvidenciasBeatriz CarvalhoNo ratings yet
- (Superpartituras - Com.br) EvidenciasDocument1 page(Superpartituras - Com.br) EvidenciasMarco A. P. AmmirabileNo ratings yet
- Chitaozinho e Xororo Evidencias PDFDocument1 pageChitaozinho e Xororo Evidencias PDFVinicius MartinsNo ratings yet
- Chitaozinho e Xororo Evidencias PDFDocument1 pageChitaozinho e Xororo Evidencias PDFPHERNANDO E KAIKENo ratings yet
- La Chocolata Azinhaga-Saxofone AltoDocument1 pageLa Chocolata Azinhaga-Saxofone Altoinesdc5No ratings yet
- Legend Barangay Boundary Barangay Road: A C A B C D E FDocument1 pageLegend Barangay Boundary Barangay Road: A C A B C D E FJoel Rodello B DugeniaNo ratings yet
- Dspa595 SchDiagram OperationDocument1 pageDspa595 SchDiagram OperationAndrew Chan YanNo ratings yet
- Est-Casa Club Aires Del Batan-E2Document1 pageEst-Casa Club Aires Del Batan-E2ISMAEL MONTIELNo ratings yet
- Bidp SDH 2022 Ar 117 DTCMCDocument1 pageBidp SDH 2022 Ar 117 DTCMCJeffNo ratings yet
- Suite Violino IDocument2 pagesSuite Violino IDaniel PivaNo ratings yet
- Oh DarlingDocument3 pagesOh DarlingJoão PedroNo ratings yet
- Kimball Et Al 2019, Figure 7Document1 pageKimball Et Al 2019, Figure 7Abby KimballNo ratings yet
- Oko Et Al 2019, Figure 8Document1 pageOko Et Al 2019, Figure 8Abby KimballNo ratings yet
- Kimball Et Al 2019, Figure 2Document1 pageKimball Et Al 2019, Figure 2Abby KimballNo ratings yet
- Kimball Et Al 2019, Figure 3Document1 pageKimball Et Al 2019, Figure 3Abby KimballNo ratings yet
- Neuwelt Et Al, Figure 2Document1 pageNeuwelt Et Al, Figure 2Abby KimballNo ratings yet
- Kimball Et Al 2019, Figure 4Document1 pageKimball Et Al 2019, Figure 4Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 5Document1 pageKimball Et Al 2018, Figure 5Abby KimballNo ratings yet
- Kimball Et Al 2019, Figure 1Document1 pageKimball Et Al 2019, Figure 1Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 8Document1 pageKimball Et Al 2018, Figure 8Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 7Document1 pageKimball Et Al 2018, Figure 7Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 1Document1 pageKimball Et Al 2018, Figure 1Abby KimballNo ratings yet
- Bullock Et Al 2018, Supplemental Figure 5Document1 pageBullock Et Al 2018, Supplemental Figure 5Abby KimballNo ratings yet
- Graph Features:: Events Sampled Per File: 9,141 Minimum Cluster Size: 5% Group Assignment: CorrectDocument1 pageGraph Features:: Events Sampled Per File: 9,141 Minimum Cluster Size: 5% Group Assignment: CorrectAbby KimballNo ratings yet
- Berger Et Al 2019, Figure 2Document1 pageBerger Et Al 2019, Figure 2Abby KimballNo ratings yet
- Kimball Et Al 2018, Figure 2Document1 pageKimball Et Al 2018, Figure 2Abby KimballNo ratings yet
- Bullock Et Al 2018, Figure 6Document1 pageBullock Et Al 2018, Figure 6Abby KimballNo ratings yet
- Hospital Pharmacy Cover LetterDocument7 pagesHospital Pharmacy Cover Letterykzdmfajd100% (2)
- Borax PDFDocument4 pagesBorax PDFZi LaNo ratings yet
- Lobes of The Brain - FrontalDocument12 pagesLobes of The Brain - FrontalLoccos SienaNo ratings yet
- Make Every Session Count PDFDocument209 pagesMake Every Session Count PDFEgylany LanyNo ratings yet
- Xylocaine Dentaland AdrenalineDocument7 pagesXylocaine Dentaland Adrenalinefawwaz FofoNo ratings yet
- CP On AnginaDocument21 pagesCP On AnginaNavpreet KaurNo ratings yet
- A Classification Tree To Assist With Routine Scoring of The Clinical Frailty ScaleDocument6 pagesA Classification Tree To Assist With Routine Scoring of The Clinical Frailty ScaleOlga GuerreroNo ratings yet
- Physical Examination by DRDocument25 pagesPhysical Examination by DRapi-3739910100% (2)
- Seht Card Hospital ListDocument33 pagesSeht Card Hospital Listlucky xenNo ratings yet
- Analytical Method Technology Transfer Is Pe GuideDocument46 pagesAnalytical Method Technology Transfer Is Pe Guidevenkat_du2000100% (1)
- Fisher 2009Document6 pagesFisher 2009Maryanne CunhaNo ratings yet
- Paul Ramesh Forensic Neuro Psychological InterviewDocument43 pagesPaul Ramesh Forensic Neuro Psychological Interviewnonam2100% (2)
- Pacemaker PamphletDocument2 pagesPacemaker Pamphletapi-355079078No ratings yet
- FSC Extra ENT and Tuning Fork January 11-Th 2018 Internet ConferenceDocument21 pagesFSC Extra ENT and Tuning Fork January 11-Th 2018 Internet Conferencemarc800No ratings yet
- Supply Chain Management in HospitalsDocument6 pagesSupply Chain Management in HospitalsAbhra100% (10)
- BNF - 76 - British - National - Formulary - Septem-357-358 PhenobarbtalDocument2 pagesBNF - 76 - British - National - Formulary - Septem-357-358 Phenobarbtalnur aisahNo ratings yet
- Prostate, Lung, Colorectal and Ovarian Cancer Screening Trial Medication Use QuestionnaireDocument3 pagesProstate, Lung, Colorectal and Ovarian Cancer Screening Trial Medication Use QuestionnaireGeede AlfonsoNo ratings yet
- Notes To Remember: For Clinical Ophthalmology MCQ ExamsDocument89 pagesNotes To Remember: For Clinical Ophthalmology MCQ ExamsMohamed Gaber100% (1)
- Bronchopulmonary Dysplasia: An Update: Anita Bhandari and Vineet BhandariDocument5 pagesBronchopulmonary Dysplasia: An Update: Anita Bhandari and Vineet BhandariPatricius Berry Harsono PutraNo ratings yet
- Cumitech 23 - Infections of The Skin and Subcutaneous Tissues PDFDocument16 pagesCumitech 23 - Infections of The Skin and Subcutaneous Tissues PDFAnjali MohanNo ratings yet
- Sucralfate PDFDocument2 pagesSucralfate PDFJoshua Christian Penggele100% (1)
- Prioritization of Health ProblemsDocument2 pagesPrioritization of Health ProblemsrorenNo ratings yet
- SL 1 MPT 23Document2 pagesSL 1 MPT 23Samidul IslamNo ratings yet
- RSI For Nurses ICUDocument107 pagesRSI For Nurses ICUAshraf HusseinNo ratings yet
- CV - DR Andrew Balyeku - June 2018 BDocument4 pagesCV - DR Andrew Balyeku - June 2018 BIga_DanNo ratings yet
- Ascorbic Acid InjectionDocument6 pagesAscorbic Acid InjectionMaria NorilynNo ratings yet
- Students With Learning Disabilities and InclusionDocument5 pagesStudents With Learning Disabilities and InclusionEditor IJTSRD100% (3)
- Asbestoswise ASEA Information SheetsDocument32 pagesAsbestoswise ASEA Information SheetsHeung Yeung CheukNo ratings yet
- Black Seed Nigella Sativa A Cure To Every DiseaseDocument6 pagesBlack Seed Nigella Sativa A Cure To Every DiseaseShahmeer KhanNo ratings yet