Professional Documents
Culture Documents
Development of A New Field-Deployable RTQPCR Workflow For COVID19 Detection
Development of A New Field-Deployable RTQPCR Workflow For COVID19 Detection
Uploaded by
Greater K. OYEJOBICopyright:
Available Formats
You might also like
- Health Education HTPDocument4 pagesHealth Education HTPizelelelsNo ratings yet
- Hematology MCQsDocument84 pagesHematology MCQsHyder Ali84% (19)
- Lithium Carbonate Drug StudyDocument3 pagesLithium Carbonate Drug Studynot your medz duranNo ratings yet
- L'Helgouach-2020-EasyCOV - LAMP Based Rapid deDocument14 pagesL'Helgouach-2020-EasyCOV - LAMP Based Rapid deGerman MadrigalNo ratings yet
- Journal Pre-Proof: Journal of Clinical VirologyDocument12 pagesJournal Pre-Proof: Journal of Clinical Virologyamer khanNo ratings yet
- Evaluation of An Antigen Detection Rapid Diagnostic Test For Detection of Sars-Cov-2 in Clinical SamplesDocument9 pagesEvaluation of An Antigen Detection Rapid Diagnostic Test For Detection of Sars-Cov-2 in Clinical Samplesjytj tjrj64fdNo ratings yet
- Panbio COVID-19 Ag Rapid Test For The Diagnosis of SARS-CoV-2 - ArticuloDocument6 pagesPanbio COVID-19 Ag Rapid Test For The Diagnosis of SARS-CoV-2 - Articulo0244761No ratings yet
- Sensitive On-Site Detection of SARS-CoV-2 by ID NOW COVID-19Document12 pagesSensitive On-Site Detection of SARS-CoV-2 by ID NOW COVID-19luis.achaNo ratings yet
- 2019 New Coronavirus Nucleic Acid Detection Laboratory Construction Plan - 20200323Document15 pages2019 New Coronavirus Nucleic Acid Detection Laboratory Construction Plan - 20200323IMUNISASI DINKESNo ratings yet
- MainDocument7 pagesMainJhessica Gonzales GarciaNo ratings yet
- Approach To Assessment of New Swabs and Viral Transport Media For SARS-CoV-2 Testing-JCM.01562-20Document7 pagesApproach To Assessment of New Swabs and Viral Transport Media For SARS-CoV-2 Testing-JCM.01562-20Medtech ChuongNo ratings yet
- Lmo011 04 290Document7 pagesLmo011 04 290aobeso002No ratings yet
- Covid19 Methods PDFDocument14 pagesCovid19 Methods PDFSulieman OmarNo ratings yet
- Jurnal MPIDocument4 pagesJurnal MPIMarini TaslimaNo ratings yet
- 10 1128@JCM 00512-20Document22 pages10 1128@JCM 00512-20Luis KrNo ratings yet
- Swab Pooling For Large-Scale RT-QPCR Screening of Sars-Cov-2Document22 pagesSwab Pooling For Large-Scale RT-QPCR Screening of Sars-Cov-2popov357No ratings yet
- Quality in Diagnostic MicrobiologyDocument6 pagesQuality in Diagnostic Microbiologymicrobehunter007No ratings yet
- Multicenter Evaluation of The Panbio™ COVID-19 Rapid Antigendetection Test For The Diagnosis of SARS-CoV-2 InfectionDocument5 pagesMulticenter Evaluation of The Panbio™ COVID-19 Rapid Antigendetection Test For The Diagnosis of SARS-CoV-2 InfectionamyNo ratings yet
- JurnalDocument6 pagesJurnalDaffa AthaNo ratings yet
- Cid RT PCR Positive Over TimeDocument12 pagesCid RT PCR Positive Over TimePatrick CoNo ratings yet
- COVID-19 Testing One Size Does Not Fit AllDocument3 pagesCOVID-19 Testing One Size Does Not Fit Alliq_dianaNo ratings yet
- Molecular and Antibody Point-Of-Care Tests To Support The Screening, Diagnosis and Monitoring of COVID-19Document12 pagesMolecular and Antibody Point-Of-Care Tests To Support The Screening, Diagnosis and Monitoring of COVID-19vicndubNo ratings yet
- Biosafety Guidelines For The Use of GeneXperts As A Point-Of-Care Test (POCT)Document4 pagesBiosafety Guidelines For The Use of GeneXperts As A Point-Of-Care Test (POCT)katrinatigaNo ratings yet
- Pooling of Samples For Sars-Cov-2 Detection Using A Rapid Antigen TestDocument5 pagesPooling of Samples For Sars-Cov-2 Detection Using A Rapid Antigen TestErickson OngNo ratings yet
- The Detection Dogs Test Is More Sensitive Than Real-Time PCR in Screening For SARS-CoV-2Document7 pagesThe Detection Dogs Test Is More Sensitive Than Real-Time PCR in Screening For SARS-CoV-2Alejandro MerelesNo ratings yet
- Estudio Corona 1 Ropa41-7-E67Document5 pagesEstudio Corona 1 Ropa41-7-E67eudaldoNo ratings yet
- The Identification of The Sars Co V 2 Whole Genome Nine Cases Among Patients in Banten Province IndonesiaDocument13 pagesThe Identification of The Sars Co V 2 Whole Genome Nine Cases Among Patients in Banten Province IndonesiaTyas AnastasyaNo ratings yet
- Challenges in Laboratory Diagnosis of The NovelDocument27 pagesChallenges in Laboratory Diagnosis of The NovelRidho Al Fiqri100% (1)
- Title: Antigen Rapid Tests, Nasopharyngeal PCR and Saliva PCR To Detect Sars-Cov-2: A Prospective Comparative Clinical TrialDocument16 pagesTitle: Antigen Rapid Tests, Nasopharyngeal PCR and Saliva PCR To Detect Sars-Cov-2: A Prospective Comparative Clinical Trialorkut verificationNo ratings yet
- AbbottDocument10 pagesAbbottSisca PrimaNo ratings yet
- 108820lateral Move Molecular Assay For Horse StranglesDocument3 pages108820lateral Move Molecular Assay For Horse StranglesneriktjpcoNo ratings yet
- 1 s2.0 S0033350620301141 MainDocument3 pages1 s2.0 S0033350620301141 MainFedrik Monte Kristo LimbongNo ratings yet
- Real-Time RT-PCR For COVID-19 Diagnosis: Challenges and ProspectsDocument3 pagesReal-Time RT-PCR For COVID-19 Diagnosis: Challenges and ProspectsAnthony StackNo ratings yet
- CT Is Not EnoughDocument4 pagesCT Is Not Enougharhur9719No ratings yet
- Supardi, 2021Document15 pagesSupardi, 2021Khumaira SantaNo ratings yet
- Understanding Antigen Tests and Results ENG FinalDocument4 pagesUnderstanding Antigen Tests and Results ENG FinalAna CatarinaNo ratings yet
- JCM 01713-20Document13 pagesJCM 01713-20Yogiswara KarangNo ratings yet
- Genes 11 00664 v2Document13 pagesGenes 11 00664 v2Monika GonzalezNo ratings yet
- Assessing A Chip Based Rapid RTPCR Test For Sars Cov-2 Detection (Truenat Assay) : A Diagnostic Accuracy StudyDocument6 pagesAssessing A Chip Based Rapid RTPCR Test For Sars Cov-2 Detection (Truenat Assay) : A Diagnostic Accuracy StudySumesh Shreekhanda ShresthaNo ratings yet
- Serologic Responses To Sars-Cov-2 Infection Among Hospital Staff With Mild Disease in Eastern FranceDocument21 pagesSerologic Responses To Sars-Cov-2 Infection Among Hospital Staff With Mild Disease in Eastern FranceAssia AouachriaNo ratings yet
- Development of A Novel Loop-Mediated Isothermal Amplification Method For The Rapid Detection of Monkeypox Virus InfectionsDocument12 pagesDevelopment of A Novel Loop-Mediated Isothermal Amplification Method For The Rapid Detection of Monkeypox Virus InfectionsV K kasaragodNo ratings yet
- Predicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 From Diagnostic SamplesDocument4 pagesPredicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 From Diagnostic SamplesmeNo ratings yet
- Up Date Diagnosis Laboratorium Pada Pedoman: Pencegahan Dan Pengendalian COVID-19 Revisi Ke-5Document65 pagesUp Date Diagnosis Laboratorium Pada Pedoman: Pencegahan Dan Pengendalian COVID-19 Revisi Ke-5Muhammad NurfiantoroNo ratings yet
- Predicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 From Diagnostic SamplesDocument4 pagesPredicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 From Diagnostic SamplesArum Rahmawatie VirgieNo ratings yet
- Jurnal 2Document10 pagesJurnal 2GFORCE .ID.No ratings yet
- ! Rapid Detection of Novel Coronavirus (COVID-19) by Reverse Transcription-Loop-Mediated Isothermal AmplificationDocument17 pages! Rapid Detection of Novel Coronavirus (COVID-19) by Reverse Transcription-Loop-Mediated Isothermal AmplificationPetr CiglerNo ratings yet
- Presence of Mismatches Between Diagnostic PCR Assays and Coronavirus Sars-Cov-2 GenomeDocument17 pagesPresence of Mismatches Between Diagnostic PCR Assays and Coronavirus Sars-Cov-2 GenomeYulaimy BatistaNo ratings yet
- Journal of Clinical Microbiology-2020-Chan-JCM.00310-20.full PDFDocument33 pagesJournal of Clinical Microbiology-2020-Chan-JCM.00310-20.full PDFDwi Ratna KarimNo ratings yet
- Multiplex Detection and Dynamics of IgG Antibodies To SARS-CoV2 and The Highly Pathogenic Human Coronaviruses SARS-CoV and MERS-CoVDocument7 pagesMultiplex Detection and Dynamics of IgG Antibodies To SARS-CoV2 and The Highly Pathogenic Human Coronaviruses SARS-CoV and MERS-CoVamyNo ratings yet
- 2020 Article 335 PDFDocument4 pages2020 Article 335 PDFFellita Ratri ANo ratings yet
- Makalah Biologi MolekulerDocument11 pagesMakalah Biologi MolekulerSarah PanjaitanNo ratings yet
- OASH COVID 19 Guidance Testing PlatformsDocument3 pagesOASH COVID 19 Guidance Testing PlatformsNurul PalesseiNo ratings yet
- Journal of Clinical Microbiology-2020-Chan-JCM.00310-20.full PDFDocument33 pagesJournal of Clinical Microbiology-2020-Chan-JCM.00310-20.full PDFdavid zhangNo ratings yet
- Ijerph-18-11451 Meta Analysis StudyDocument14 pagesIjerph-18-11451 Meta Analysis StudySumit BediNo ratings yet
- Different Longitudinal Patterns of Nucleic Acid and Serology Testing Results Based On Disease Severity of COVID-19 PatientsDocument5 pagesDifferent Longitudinal Patterns of Nucleic Acid and Serology Testing Results Based On Disease Severity of COVID-19 PatientsagboxNo ratings yet
- Towards Detection of SARS-CoV-2 RNADocument8 pagesTowards Detection of SARS-CoV-2 RNARuben MonroyNo ratings yet
- Correspondence To: Professor Changxin Shen Department of Blood TransfusionDocument13 pagesCorrespondence To: Professor Changxin Shen Department of Blood TransfusionAlfa RidziNo ratings yet
- Lynx PublicationDocument9 pagesLynx PublicationSwarnendraNo ratings yet
- Performance Verification of Anti-Sars-Cov-2-Specific Antibody Detection by Using Four Chemiluminescence Immunoassay SystemsDocument6 pagesPerformance Verification of Anti-Sars-Cov-2-Specific Antibody Detection by Using Four Chemiluminescence Immunoassay SystemsadnanNo ratings yet
- A Role For Biofoundries in Rapid Development and Validation of Automated Sars-Cov-2 Clinical DiagnosticsDocument11 pagesA Role For Biofoundries in Rapid Development and Validation of Automated Sars-Cov-2 Clinical DiagnosticsRicardoPeixotoNo ratings yet
- Validasi Rapid AntigenDocument6 pagesValidasi Rapid Antigengaki telitiNo ratings yet
- Artigo Cientifico Barreto2020 Article Diagnosingthenovelsars Cov 2bypdfDocument10 pagesArtigo Cientifico Barreto2020 Article Diagnosingthenovelsars Cov 2bypdfOtaku GamerChanNo ratings yet
- SARS-CoV-2 Viral Outbreak Investigation: Laboratory Perspective: Clinical Updates in COVID-19From EverandSARS-CoV-2 Viral Outbreak Investigation: Laboratory Perspective: Clinical Updates in COVID-19Rating: 3 out of 5 stars3/5 (1)
- Athlete's Heart: Dr. Arzalan BaigDocument59 pagesAthlete's Heart: Dr. Arzalan BaigArzalan BaigNo ratings yet
- TRẮC NGHIỆM AVCN PHARMACYDocument91 pagesTRẮC NGHIỆM AVCN PHARMACYHoang Pham ThaiNo ratings yet
- Fisiologia Do EnvelhecimentoDocument2 pagesFisiologia Do EnvelhecimentoLissandro DannyNo ratings yet
- Physical Examination FormatDocument6 pagesPhysical Examination Formatvethadhas vin100% (1)
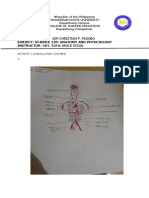
- Student'S Name: Kim Christian P. Pagobo Subject: Science 120: Anatomy and Physciology Instructor: Mrs. Roma Grace DizonDocument9 pagesStudent'S Name: Kim Christian P. Pagobo Subject: Science 120: Anatomy and Physciology Instructor: Mrs. Roma Grace Dizonkim christianNo ratings yet
- Coronary Artery Diseases - Ilham PDocument43 pagesCoronary Artery Diseases - Ilham PKeputrian FKUPNo ratings yet
- Patient Classification & AssignmentDocument37 pagesPatient Classification & AssignmentAkhila UsNo ratings yet
- J.H. Cerilles State College: Western Mindanao State UniversityDocument6 pagesJ.H. Cerilles State College: Western Mindanao State UniversityVanessa PalomaNo ratings yet
- Nclex Study GuideDocument37 pagesNclex Study GuideBlas Christian Tancongco ChavesNo ratings yet
- Fermented Products With Probiotic Qualities - Casei & Bifidobakterije PDFDocument6 pagesFermented Products With Probiotic Qualities - Casei & Bifidobakterije PDFmilu1312No ratings yet
- Bigay-Panghulan - Legal Opinion - Legal Forms - First ActivityDocument3 pagesBigay-Panghulan - Legal Opinion - Legal Forms - First ActivityApril Ann Sapinoso Bigay-PanghulanNo ratings yet
- UHSAA Preparticipation FormDocument4 pagesUHSAA Preparticipation FormThe Salt Lake TribuneNo ratings yet
- Functional Constipation: A Case Report: EpartmentDocument5 pagesFunctional Constipation: A Case Report: EpartmentArielIzaSuquilloNo ratings yet
- Univestin+ Glucosmine NewDocument26 pagesUnivestin+ Glucosmine NewAshishNo ratings yet
- Finalised ILERA EKO DIASPORA PLAN PRODUCT PAPER 1Document19 pagesFinalised ILERA EKO DIASPORA PLAN PRODUCT PAPER 1dibransonsNo ratings yet
- Donning Doffing Spotter Check List v1Document1 pageDonning Doffing Spotter Check List v1Rudyard Paul AmistadNo ratings yet
- Lab 2,3Document18 pagesLab 2,3Nosheela KhalidNo ratings yet
- New Patient Nutrition Assessment Form: (Person 1)Document12 pagesNew Patient Nutrition Assessment Form: (Person 1)Ayesha KhanNo ratings yet
- Krauses Food and The Nutrition Care Process 14Th Edition Mahan Test Bank Full Chapter PDFDocument28 pagesKrauses Food and The Nutrition Care Process 14Th Edition Mahan Test Bank Full Chapter PDFphualexandrad39100% (12)
- Blaylock Hypertension 13Document16 pagesBlaylock Hypertension 13mohanp71No ratings yet
- Bloodbld 2019003808 CDocument12 pagesBloodbld 2019003808 CBillal BenhaddadNo ratings yet
- OSCES 5e MMSDocument125 pagesOSCES 5e MMSFlórian Jefferson100% (1)
- Vinblastine (Velban®) : Things That May Occur During or Within Hours of TreatmentDocument2 pagesVinblastine (Velban®) : Things That May Occur During or Within Hours of TreatmentVette Angelikka Dela CruzNo ratings yet
- Rehan Aftab Hamdulay, Himanshu Girish Sharma, Bombay College of Pharmacy, Santacruz (East), Mumbai - 400098Document1 pageRehan Aftab Hamdulay, Himanshu Girish Sharma, Bombay College of Pharmacy, Santacruz (East), Mumbai - 400098Himanshu SharmaNo ratings yet
- Inguinal Hernia Case FileDocument2 pagesInguinal Hernia Case Filehttps://medical-phd.blogspot.comNo ratings yet
- Pranav SharmaDocument11 pagesPranav SharmaPranav SharmaNo ratings yet
- Kapalbhati PDFDocument2 pagesKapalbhati PDFRahul MishraNo ratings yet
Development of A New Field-Deployable RTQPCR Workflow For COVID19 Detection
Development of A New Field-Deployable RTQPCR Workflow For COVID19 Detection
Uploaded by
Greater K. OYEJOBIOriginal Description:
Original Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Development of A New Field-Deployable RTQPCR Workflow For COVID19 Detection
Development of A New Field-Deployable RTQPCR Workflow For COVID19 Detection
Uploaded by
Greater K. OYEJOBICopyright:
Available Formats
Life Research doi: 10.
53388/life2021-0525-110
ARTICLE
Development of a new field-deployable RT-qPCR workflow for COVID-
19 detection
Raphael Nyaruaba1, 2, 3, Bo Zhang1, Caroline Muema1, 2, 3, Elishiba Muturi1, 2, 3, Greater K. Oyejobi1, 2, Jin Xiong1, Bei Li1, Zheng-Li Shi1,
Caroline Mwaliko1, 2, 3, Jun-Ping Yu1, Xiao-Hong Li1, Ya-Nan Zhang1, Hong-Ping Wei1, 3 *
Key Laboratory of Special Pathogens and Biosafety, Center for Biosafety Mega-Science, Wuhan Institute of Virology, Chinese
1
Academy of Sciences, Wuhan, Hubei, China. 2International College, University of Chinese Academy of Sciences, Beijing, China. 3Sino-
Africa Joint Research Center, Nairobi, Kenya.
*Corresponding to: Hong-Ping Wei. Key Laboratory of Special Pathogens and Biosafety, Center for Biosafety Mega-Science, Wuhan
Institute of Virology, Chinese Academy of Sciences, No. 44, Bayi Road, Wuhan 430071, China. E-mail: hpwei@wh.iov.cn.
Abstract
Background: Outbreaks of coronavirus disease 2019 (COVID-19) have been recorded in different countries across
the globe. The virus is highly contagious, hence early detection, isolation, and quarantine of infected patients will
play an important role in containing the viral spread. Diagnosis in a mobile lab can aid to find infected patients in
time. Methods: Here, we develop a field-deployable diagnostic workflow that can reliably detect COVID-19.
Instruments used in this workflow can easily fit in a mobile cabin hospital and also be installed in the community.
Different steps from sample inactivation to detection were optimized to find the fastest steps and portable instruments
in the detection of COVID-19. Each step was compared to that of the normal laboratory diagnosis setup. Results:
From the results, our proposed workflow (80 min) was two times faster compared to that of the normal laboratory
workflow (183 min) and a maximum of 32 samples could be detected at each run. Additionally, we showed that using
1% Rewocid WK-30 could inactivate the novel coronavirus directly without affecting the overall detection results.
Comparison of our workflow using an in-house assay to that of a commercially acquired assay produced highly
reliable results. From the 250 hospital samples tested, there was a high concordance 247/250 (98.8%) between the
two assays. The in-house assay sensitivity and specificity were 116/116 (100%) and 131/134 (97.8%) compared to
that of the commercial assay. Conclusion: Based on these results, we believe that our workflow is fast, reliable,
adaptable and most importantly, field-deployable.
Key words: COVID-19, SARS-CoV-2, Field work, Community, Diagnosis, Rapid detection, RT-qPCR
Competing interests:
The authors declare that they have no conflict of interest.
Acknowledgement:
All the researchers acknowledge the help and support provided by all students and staff at the Key Laboratory of
Special Pathogens and Biosafety, Center for Biosafety Mega-Science, Wuhan Institute of Virology, Chinese
Academy of Sciences.
Abbreviations:
COVID-19, coronavirus disease 2019; SARS-CoV-2, serve acute respiratory syndrome coronavirus 2; PCR,
polymerase chain reaction; RT-qPCR, real-time reverse transcription-quantitative polymerase chain reaction;
VTM, viral transport medium; FBS, fetal bovine serum; DMEM, Dulbecco’s Modified Eagle Medium; POC, point
of care; Ct, cycle threshold.
Citation:
Nyaruaba R, Zhang B, Muema C, et al. Development of a new field-deployable RT-qPCR workflow for COVID-
19 detection. Life Res. 2021;4(3):24. doi: 10.53388/life2021-0525-110.
Executive Editor: Shan-Shan Lin.
Submitted: 25 May 2021, Accepted: 03 July 2021, Online: 24 July 2021
© 2021 By Authors. Published by TMR Publishing Group Limited. This is an open access article under the CC-
BY license (http://creativecommons.org/licenses/BY/4.0/).
Submit a manuscript: https://www.tmrjournals.com/lr 1
doi: 10.53388/life2021-0525-110 ARTICLE
practical workflow that is fast and can be easily adapted
Background in different facilities. The workflow also includes a
field-able approach for the diagnosis in the community
Coronavirus disease 2019 (COVID-19) is caused by the and mobile cabin hospitals set-up. The workflow was
serve acute respiratory syndrome coronavirus 2 (SARS- compared to the standard workflow using the
CoV-2). In December 2019, the first cases of human conventional RT-qPCR diagnosis of the ongoing
infection with the COVID-19 were identified in Wuhan, coronavirus outbreak.
China [1–3]. Most of the cases were linked to a local
seafood market in Wuhan which is believed to be the Methods
source of the SARS-CoV-2 virus outbreak [2–5]. Since
its identification, scientists have characterized the virus Sample collection
[3, 6, 7] and reported clinical symptoms associated with Wuhan Institute of Virology CAS is one of the
COVID-19 [2, 3, 8, 9]. Controlling the virus has been a authorized labs approved by the Centers for Disease
top priority in areas affected both in China and across Control of Wuhan city for detecting COVID-19 in
the globe. As of April 8th, 2020, the World Health clinical samples. All the samples were handled and
Organization daily situation report on the novel deactivated first in a biosafety level 2 laboratory with
coronavirus disease recorded a total of 1,403,622 personal protection equipment for biosafety level 3 lab
(87,279 new) confirmed cases and 85,495 (8,902 new) following the guidelines for detecting nucleic acid of
total deaths resulting from COVID-19 globally [10]. COVID-19 in clinical samples. Research on developing
Cases of the epidemic outbreak have also been reported new diagnostic techniques for COVID-19 using clinical
in nearly all countries across the globe [10, 11]. With samples has been approved by the ethical committee of
the daily rise in number of deaths, suspected, and Wuhan Institute of Virology (2020FCA001). Human
confirmed cases across the globe, methods towards samples in the form of oral swabs were collected from
rapid diagnosis and detection of the virus disease are of various health facilities across Wuhan and transported
key importance in fighting the pandemic. to our laboratory for detection. The swab samples were
From the beginning of the pandemic, different suspended in tubes containing viral transport medium
diagnostic approaches have been used in an attempt to (VTM) prior to transportation. The VTM constituted
tame and understand the spread of infections resulting Hank’s balanced salt solution at pH 7.4 containing BSA
from SARS-CoV-2 [1, 3, 8, 12–15]. Among these (1%), amphotericin (15 μg/mL), penicillin G (100
approaches includes the development of specific viral units/mL), and streptomycin (50 μg/mL). Upon receipt
nucleic acid assays that use real-time reverse at Wuhan Institute of Virology, Zhengdian, the samples
transcription-quantitative polymerase chain reaction were carried in locked containers and moved to the
(RT-qPCR) to diagnose new cases of COVID-19 [1, 3, biosafety level 2 laboratory. In the biosafety level 2
16–18]. RT-qPCR applications in diagnostics however llaboratory, the samples were inactivated by heating at
are not a new concept. Since its advent, RT-qPCR has 56 ℃ for 30 min. The samples were then left to cool at
become a well-established technique for the diagnosis 4 ℃ before immediate processing or stored at the same
of various microorganisms owing to its many temperature until processing.
advantages including rapidity, sensitivity, ease of
application, and scalability [19]. Many companies have Sample processing
come up with different qPCR instruments that are lab- To obtain the fastest workflow, we developed a sample
based and also field deployable. In the past coronavirus matrix of 3 categories for spiking and testing the whole
outbreaks [20], RT-qPCR has also been used workflow from sample collection to detection. These
extensively to help in the identification of different categories included: (1) swabs collected from hospitals
cases and help in the diagnosis pipeline of the virus in VTM tubes; (2) swabs spiked in RNase free water;
outbreaks [21–23]. However, in these outbreaks and (3) swabs spiked with already tested positive
including the ongoing COVID-19, no research has tried samples. The time and data for all the above sample
to describe an optimized system and workflow suitable categories were collected in each of the following
for point of care testing, field deployment, and one that processing steps.
can easily fit in a mobile cabin hospital to detect and Sample inactivation. The normal sample inactivation
diagnose the viral disease. The development of efficient procedure involved heating swab samples in VTM tubes
and quicker methods for the detection of viral nucleic at 56 ℃ for 30 min and letting potential aerosols settle
acids is said to play an important role in fighting the for an extra 10 min at 4 ℃ before extraction. To find a
ongoing coronavirus outbreak [12]. Additionally, the faster way suitable for diagnosis at field conditions, we
development of a workflow that can fit in a mobile tried an approach of using the biocidal Rewocid WK-30
hospital can guarantee comprehensive healthcare to (Evonik Industries, Shanghai HonenesteverCo. Ltd.,
anybody, anytime and anywhere [24–27]. Shanghai, China) here referred to as WK-30 at a
In light of these limitations, we sought to develop a concentration of 1% to directly inactivate the virus
2 Submit a manuscript: https://www.tmrjournals.com/lr
Life Research doi: 10.53388/life2021-0525-110
without heating. The virucidal and bactericidal activity Different approaches were also considered for sample
of WK-30 against COVID-19 and different bacterial detection.
strains was also tested. RT-qPCR quantification instruments. To find a
Virucidal activity tests. For virucidal testing, three suitable qPCR instrument that is field-deployable and
tests were done independently by mixing 2% WK-30, suitable for POC testing, we used a portable MyGo Pro
1.6 × 106 PFU/mL SARS-CoV-2, and Dulbecco’s Real-time PCR instrument readily available in our
Modified Eagle Medium (DMEM) supplemented with laboratory to optimize our workflow. The machine
2% fetal bovine serum (FBS) according to the following detection time was compared to that of the routinely
ratios: (1) 50 µL 2% WK-30 + 50 µL 2% FBS DMEM; used CFX-96 Touch™ Real-time PCR (CFX-96, Bio-
(2) 50 µL SARS-CoV-2 + 50 µL 2% FBS DMEM; and Rad Laboratories, Hercules, CA, USA) instrument
(3) 50 µL 2% WK-30 + 50 µL SARS-CoV-2. The suitable for laboratory benchtop diagnosis.
prepared mixtures were left to stand at room RT-qPCR assay composition and reagent
temperature for 1 min and then passed through a sieving preparation. Two assays were used for time
column to collect the first 100 µL elute. The elutes were comparison and workflow tests. The two assays
then inoculated into the Vero E6 cell and incubated for comprised of a commercially acquired kit for RT-qPCR
48 hours to observe the cytopathic effects and measure and an in-house assay developed using locally available
the viral RNA concentration using the RT-qPCR kit. reagents to perform RT-qPCR. The commercially
Bactericidal activity tests. Five bacterial strains acquired kit was labelled New Coronavirus 2019-nCoV
(Staphylococcus aureus, Escherichia coli, Nucleic Acid Detection Kit (Fluorescence PCR method)
Pseudomonas aeuriginosa, Enterococcus faecalis, and produced by the Zhongshan Daan Gene Company,
Acinetobacter baumannii) were used in testing the Guanzhou, China. The kit is a one-step RT-qPCR kit
bactericidal effects of WK-30 at two concentrations. designed to target the open reading frame 1ab and
The bacterial strains were cultured in Luria broth for 24 nucleocapsid protein genes of the novel coronavirus
hours before being transferred to different falcon tubes. sequence. The nucleocapsid protein gene was labelled
The original colony forming units of each bacteria was with a FAM reporter dye while the open reading frame
then determined before adding WK-30. After colony 1ab gene was labelled with a VIC reported dye. The kit
forming units determination, each bacteria was mixed also had an endogenous internal control dye labelled
with WK-30 to final concentrations of 0.5% and 1% Cy5. The 25 µL reaction mixture consisted of 20 µL of
WK-30. After mixing each bacteria with WK-30, the freshly prepared mix and a 5 µL RNA template. The
tubes were left to sit for 5, 10, 20, and 30 minutes at one-step RT-qPCR protocol was run using the Bio-
room temperature. 10 µL was subsequently collected at Rad’s CFX-96 instrument under the following
each time point and plated onto solid LB media plates. conditions: 50 ℃ for 15 min, 95 ℃ for 15 min, followed
All the plates were then incubated at 37 ℃ for 24 hours. by 45 cycles of 95 ℃ for 15 sec and reading at 55 ℃ for
Post incubation, the plates were checked to determine 45 sec respectively.
the presence of any colonies. The in-house assay was a one-step RT-qPCR assay
RNA extraction. Four methods of RNA extraction composed of two sets of primers and probes. A set of
were evaluated to get the fastest method of extraction. primers was designed to target the receptor binding
The goal was to find a portable extraction method that domain of the novel corona virus sequence while the
can easily be used in the field setup. The four methods other set was designed to detect an endogenous internal
included using the Qiagen Viral RNA extraction kit control. The primers and probe targeting the receptor
(spin protocol and vacuum protocol), QIAxtractror binding domain sequence include: forward primer
Automated extraction, and Purifier™ Modesty CTCAAGTGTCTGTGGATCACG; reverse primer
automated RNA extraction. Each time the extraction CCTGTGCCTGTTAAACCATTG; and probe 5’-
methods were used, the manufacturers’ FAM-ACAGCATCAGTAGTGTCAGCAATGTCTC-
recommendations were followed. To determine if WK- BHQ1-3’. The internal control primers and probe
30 had an effect in the extraction process, seven hospital sequence: forward primer
samples that had previously tested positive by RT- AGATTTGGACCTGCGAGCG; reverse primer
qPCR were treated with WK-30 to a final concentration GAGCGGCTGTCTCCACAAGT; and probe 5’-FAM-
of 1% and the RNA extracted. The same samples were TTCTGACCTGAAGGCTCTGCGCG-BHQ1-3’.
also diluted using RNase-free water as a control to a Before addition, all the primer and probe concentrations
final concentration of 1% for testing the normal were adjusted to 10 µM. The composition of the in-
extraction procedure. The resultant cycle threshold (Ct) house assay included 0.8 µL of each primer, 1 µL of
values of the two tests were determined using the each probe, 2 µL 5× PrimeScript RT Master, 1.6 µL
portable MyGo Pro real-time PCR instrument (IT-IS dNTP, 0.5 µL Taq polymerase, 3.7 µL RNase free water,
Life Science Ltd., Mahon, Cork, Ireland). and 5 µL RNA template to a final volume of 20 µL. All
the in-house one-step RT-qPCR reaction mixtures were
Sample detection then amplified using the MyGo Pro Real-time PCR
Submit a manuscript: https://www.tmrjournals.com/lr 3
doi: 10.53388/life2021-0525-110 ARTICLE
instrument under the following conditions: 50 ℃ for 10 quantified at time 0 hours and 48 hours of incubation.
min, 95 ℃ for 10 sec, followed by 40 cycles of 95 ℃ The Ct greatly dropped from Ct 19.51 at time 0 hours to
for 5 sec and reading at 60 ℃ for 30 sec respectively. Ct 34.46 at time 48 hours. From these results, 1% WK-
30 showed some level of protection to the cells. Also,
Assay specificity and performance the interaction of 1% WK-30 with SARS-CoV-2 for 1
For specificity testing of the in-house assay, an in-silico min at room temperature can kill all of the SARS-CoV-
test using the basic local alignment search tool available 2.
online from National Center for Biotechnology For the bactericidal effects, five bacteria were used.
Information was used. To test whether there will be a From the results (Table 1), all the colonies were reduced
remarkable difference between the in-house assay and with increased exposure time to WK-30. E. coli, E.
the commercial kit, 250 hospital samples were extracted faecalis, and A. bauminii were completely inhibited by
and used for detection. The commercially available kit 0.5% WK-30 at all-time points. However, S. aureus and
was used as a reference kit for validating the workflow P. aeruginosa showed some level of resistance to 0.5%
of the in-house assay. All the commercial kit tests were WK-30. S. aureus was the most resistant bacteria
done using the Bio-Rad’s CFX-96 real-time PCR because even after 30 min of exposure, some colonies
instrument while all the in-house assay tests were done still grew post incubation. Of note, 1% WK-30 however
using the portable MyGo Pro real-time PCR instrument. was able to kill all the bacteria as no colonies were
observed post incubation. 1% WK-30 had a maximum
Data analysis bactericidal effect to all the bacterial strains. From these
All RT-qPCR Ct values were generated using the results, 1% WK-30 was both bactericidal and virucidal
specific PCR instruments and read on the hence suitable for use in the field as it requires no extra
accompanying software. Analytical comparisons for sample treatments and reduces the time needed for
clinical samples were performed online using the inactivation and sample transportation.
MedCalc statistical software
(https://www.medcalc.org/calc/diagnostic_test.php). RNA extraction
Bacterial colony forming units was determined by Different methods were compared to find the fastest,
counting visible colonies on the respective culture portable and easy-to-use RNA extraction mechanism in
plates. a field setup. Factors including time, the number of
sample handling steps, and any additional instrument
Results needed were also determined as shown in Table 2. The
comparison revealed that using the Purifier™ Modesty
Sample inactivation instrument was faster and needed fewer sample
Two approaches were used to determine the fastest handling steps compared to other methods. The
sample inactivation procedure that may be suitable for instrument was easily portable and fitted with an
field applications and POC testing. The normal routine ultraviolet lamp that can help in decontamination and
sample inactivation procedure which involved heating ensuring a clean environment. Due to these advantages,
the swab samples suspended in VTM tubes, took up to the instrument was found suitable for field deployment.
40 minutes to inactivate the sample. The inactivation Since 1% WK-30 was found to be virucidal, we tested
process also needed a heating source and cooling source if the solution would have an effect in the extraction
for inactivation. Hence, this was not readily suitable for process when using the Purifier™ Modesty instrument.
field deployment. Therefore, a direct approach using the The result of addition of 1% WK-30 to the virus was
biocidal surfactant WK-30 was explored. The tested in comparison of using 1% RNase free water in
surfactants’ ability to kill different bacteria and SARS- the same sample to replace WK-30. The results as
CoV-2 was established. shown in Table 3 and Figure 2 proved that 1% WK-30
The virucidal effect of 1% WK-30 against SARS- had no remarkable effect in the extraction process and
CoV-2 was tested in vitro using Vero E6 cells as shown the results were highly comparable.
in Figure 1. From the results, it was clear that 1% WK-
30 had no effect on Vero E6 cells even after 48 hours of Detection time
incubation as shown in Figure 1A. However, incubating To find the shortest time that could detect samples faster
the cells with 1.6 × 106 PFU/mL SARS-CoV-2 proved and reliably, we modified our protocol to run for a total
to be lethal. All the cells were dead after 48 hours time of 47 min 17 sec using the MyGo Pro instrument.
incubation with clear cytopathic effects as shown in This time was 64 min 2 sec shorter compared to that of
Figure 1B. Most of the cells were alive with no clear the commercially acquired New Coronavirus 2019-
cytopathic effects after 48 hours incubation with both 1% nCoV Nucleic Acid Detection Kit which runs for 1 hour
WK-30 and 1.6 × 106 PFU/ml SARS-CoV-2 as shown 51 min 37 sec according to the manufactures’ procedure.
in Figure 1C. Additionally, the Ct values were
4 Submit a manuscript: https://www.tmrjournals.com/lr
Life Research doi: 10.53388/life2021-0525-110
(A) (B) (C)
Figure 1 Virucidal effect of 1% WK-30 against the novel coronavirus. (A) Most of the cells are alive and with
no clear cytopathic effects after 48 hours incubation with 50 µL 2% WK-30 + 50 µL 2% FBS DMEM. (B) All the
vero E6 cells are dead with clear cytopathic effects after 48 hours incubation with 50 µL 2019-nCoV + 50 µL 2%
FBS DMEM. (C) Most cells are alive with no clear cytopathic effects after 48 hours incubation with 50 µL 2% WK-
30 + 50 µL 2019-nCoV. DMEM, Dulbecco’s Modified Eagle Medium.
Table 1 Bactericidal effect on different bacteria using 0.5% and 1% WK-30
CFU before S. aureuss E.colii P.aeruginosaa E.faecaliscis A.baumanniii
addition of WK30 8
addition of WK-30 3.3 × 10 5.1 × 107 7.7 × 106 4.3 × 108 3.7 × 108
Conc of WK-30 0.5% 1% 0.5% 1% 0.5% 1% 0.5% 1% 0.5% 1%
2
CFU after 5 mins TNTC 0 0 0 8 × 10 0 0 0 0 0
CFU after 10 mins TNTC 0 0 0 1 × 102 0 0 0 0 0
4
CFU after 20 mins 3.31 × 10 0 0 0 0 0 0 0 0 0
CFU after 30 mins 6.9 × 103 0 0 0 0 0 0 0 0 0
CFU, colony forming units; TNTC, too numerous to count.
Table 2 Comparison of four extraction methods
Method/ Extraction Kit Automated Portable Manual Centrifugation Samples Time
Instrument steps steps (min)
Purifier™ Genfine Yes Yes None None 32 21
Modesty
QIAxtractor QIAamp® 96 Yes No None None 96 120
QIAcube® HT Kit
(5)
Spin protocol QIAamp Viral No Yes* Multiple All Flexible ~70
RNA Mini Kit
Vacuum QIAamp Viral Partially Yes* Multiple 2 24 ~40
protocol RNA Mini Kit
*May be portable provided there is a source of centrifugation and biosafety cabinet.
Purifier™ Modesty automated instrument used the least time to extract samples with minimal sample handling steps.
Table 3 Testing the effect of 1%WK-30 in extraction
Sample Ct value of samples spiked with
1% WK-30 1% RNase free water
Submit a manuscript: https://www.tmrjournals.com/lr 5
doi: 10.53388/life2021-0525-110 ARTICLE
20200126-06 32.604/32.606 32.273/32.525
20200126-07 23.253/23.198 23.142/23.146
20200126-09 34.444/35.174 34.392/35.655
20200126-17 31.415/31.105 31.967/31.276
20200126-23 28.914/28.764 29.02/28.837
20200126-25 26.719/26.593 28.186/28.277
20200126-26 29.707/29.928 30.602/30.807
From the results of the seven already tested positive samples, it was clear that 1%WK-30 had no effect in the
extraction process and overall result.
Figure 2 Representative RT-qPCR results after extraction with 1% WK-30 (black curve) and 1% RNase free
water (purple). From the results, 1% WK-30 had no effect on the extraction process and overall result.
Assay sensitivity and specificity assay as 119/250 (47.6%) hospital samples tested
An in-silico probe and primer test using the basic local positive for the novel coronavirus using the in-house
alignment search tool available online from NCBI assay. Of note, the concordance between the two assays
resulted in 100% specificity of the in-house primers and was high 247/250 (98.8%). However, three samples that
probes in the detection of the novel coronavirus. The tested negative by the commercial assay tested positive
performance of the whole workflow including reagents by the in-house assay resulting in a specificity of
was tested using 250 patient samples. The result of this 97.76%. All the samples that tested positive by the
test is summarized in Table 4. From the results, the commercial assay also tested positive by the in-house
novel coronavirus was detected in 116/250 (46.4%) assay 116/166 (100% sensitivity) indicating that the in-
hospital samples using the commercial kit. The in-house house assay was greatly comparable to the commercial
assay was a little more sensitive than the commercial assay with improved sensitivity.
Table 4 Agreement of the commercial lab assay and the field-deployable assay for detection of swabs from
suspected patients with COVID-19 infection
N = 250 Commercial assay Sensitivity Specificity
Positive Negative (95% CI) (95% CI)
6 Submit a manuscript: https://www.tmrjournals.com/lr
Life Research doi: 10.53388/life2021-0525-110
Field assay Positive 116 3 100% 97.76%
Negative 0 131 (96.87%–100%) (93.60%–99.54%)
Calculated online by MEDCALC (https://www.medcalc.org/calc/diagnostic_test.php). CI, confidence interval.
Discussion available extraction methods and instruments tested in
this workflow, the Purifier™ Modesty instrument was
Since the discovery of the first case of COVID-19 in the fastest and most portable. The instrument was also
Wuhan, China [1–3], scientists have been working hard fitted with an ultraviolet light source to ensure the
to diagnose the disease. Efforts geared towards the cleanliness of the extraction environment pre and post
diagnosis of COVID-19 have been patient centered and extraction. Up to 32 samples could be extracted in a
applied in hospital set-ups. However, with the rising minimal time of about 21 minutes. This was highly
numbers of cases across the globe [11, 28, 29], more scalable as 32 patients could be served in a single run of
efforts should be explored especially in community extraction.
diagnostics. There may be a number of cases being To make the workflow complete, we coupled the
missed from patients in the community hence field work workflow with a fast detection protocol using a portable
tests may prove beneficial in the treatment and MyGo Pro real-time PCR instrument for detection. The
diagnosis of COVID-19. A field based diagnostic instrument was highly compatible with the Purifier™
approach has not yet been explored. In this article, we Modesty nucleic acid extractor as it could also process
used locally available instruments and reagents to a maximum of 32 samples. The fast RT-qPCR protocol
develop a rapid RT-qPCR diagnostic workflow that is ensured samples were detected within 47 min 17 sec.
not only suitable for POC but also field deployable. In This detection time was faster and results were highly
the field setup, a rapid direct sample inactivation comparable with that of the commercial assay. The
method that needs no additional instruments could be of MyGo Pro real-time PCR instrument could be run with
great importance [27]. To develop such a method, we a USB drive. This meant that multiple instruments could
explored the biocidal effects of Rewocid WK 30 [30]. be used and run with only a single computer using
We tested if the surfactant had the potential of killing multiple USB programmed drives. The multiple options
both bacteria and viruses. In both cases, 1% WK-30 was for running the PCR instrument gave room for scaling.
shown to be virucidal to SARS-CoV-2 and also Portable instruments are important for onsite diagnosis
bactericidal to different bacteria in vitro. The virucidal and commutability [31]. All the instruments used in the
effects of 1% WK-30 also had no effect on the optimization of the proposed workflow were highly
extraction process. So far, no literature has cited or tried portable (Figure 3). This meant that the workflow
to explore the biocidal capabilities of Rewocid WK 30. instruments could easily be fitted and adapted in a
The ability of 1% WK-30 to inactivate and kill the novel mobile hospital before deployment to the community to
coronavirus at room temperature within a minute of detect and diagnose COVID-19. The complete
exposure makes it a suitable reagent for direct sample workflow from sample inactivation to detection was
inactivation in the field set-up. However, when handling approximated to be no more than 80 minutes using the
samples, one should also take care to observe biosafety field deployable workflow. This workflow was two
and wear the appropriate personal protective equipment. times faster compared to that of the normal workflow in
Once the sample has been inactivated, the sample the laboratory setup (183 min 9 sec). Additionally, after
should proceed directly to extraction of the viral nucleic the first detection of approximately 80 minutes, the
acid necessary for detection. A fast, portable automated proceeding samples can be extracted and detected
extractor with minimal sample handling steps will be within an hour. As the first samples are running, a new
suitable for field extraction experiments. Automation batch of samples may be loaded into the extractor which
with minimal sample handling steps ensures safety to will run for 21 min, the extra 26 minutes before the RT-
the worker and also ensures sample integrity is kept qPCR results are read can be used for reaction mix
intact throughout extraction in the field [27] where there preparation and programming. Once the first batch
are minimal or no specialized equipment e.g. biosafety finishes, the newly extracted samples can be detected
cabinets. A portable instrument on the other hand meant immediately.
that it could fit well in a mobile hospital. Of all the
Submit a manuscript: https://www.tmrjournals.com/lr 7
doi: 10.53388/life2021-0525-110 ARTICLE
Figure 3 Schematic comparison of the proposed workflow compared to the normal lab-based detection
workflow. The proposed workflow is faster than the normal lab-based workflow. All the instruments in the proposed
workflow are portable.
This workflow is a first in exploring the potential of POC. We found that using Rewocid WK 30 as a
community diagnosis using a field deployable nucleic biocidal agent could help in the inactivation of SARS-
acid detection system for COVID-19. All the CoV-2 samples directly without the need of any extra
instruments highlighted to be used in the field set-up are equipment. This workflow was two times faster than
highly portable and can fit well in a mobile hospital. The that of the normal workflow in the laboratory set-up. We
instruments can also be powered by a car battery and be believe that the workflow described is easily adaptable
used in remote locations where electricity is not and flexible. This workflow will help in the fight against
accessible. Additionally, the workflow is flexible and the current SARS-CoV-2 pandemic and other future
can be modified depending on the available resources in outbreaks. Lastly, the workflow may be useful in
different laboratories and countries. The workflow can routine sample collection and detection in future field-
also be set up in hospitals, clinics, and public health work surveillance studies.
laboratories that cannot access electricity. We plan to
actualize this workflow in our subsequent tests using a References
mobile hospital to help in the diagnosis of COVID-19.
Actualization of this workflow ensures that patients are 1. Zhu N, Zhang D, Wang W, et al. A novel
treated anywhere and at any time [25, 31]. This will also coronavirus from patients with pneumonia in
help in reducing the large number of patients that visit China, 2019. N Engl J Med. 2020;382(8):727–733.
hospitals. Patient treatment at community level Available: 10.1056/NEJMoa2001017;
decreases their risks of getting infected when visiting https://doi.org/10.1056/NEJMoa2001017
the hospitals. Different researchers and laboratories can 2. Chan JFW, Yuan S, Kok KH, et al. A familial
use our workflow and modify it if needed to best fit their cluster of pneumonia associated with the 2019
laboratory. novel coronavirus indicating person-to-person
transmission: a study of a family cluster. Lancet.
Conclusion 2020;395(10223):514–523.
Available: 10.1016/S0140-6736(20)30154-9;
Portable instruments that can be used in the extraction https://doi.org/10.1016/S0140-6736(20)30154-9
and detection of nucleic acids already exist. We used 3. Zhou P, Yang XL, Wang XG, et al. A pneumonia
locally available instruments and reagents to develop a outbreak associated with a new coronavirus of
new field-deployable workflow that can also be used at probable bat origin. Nature. 2020;579(7798): 270–
8 Submit a manuscript: https://www.tmrjournals.com/lr
Life Research doi: 10.53388/life2021-0525-110
273. Available: 10.1038/s41586-020-2012-7; 14. Nyaruaba R, Li C, Mwaliko C, et al. Developing
https://doi.org/10.1038/s41586-020-2012-7 multiplex ddPCR assays for SARS-CoV-2
4. Tang X, Wu C, Li X, et al. On the origin and detection based on probe mix and amplitude based
continuing evolution of SARS-CoV-2. Natl Sci Rev. multiplexing. Expert Rev Mol Diagn. 2021;
2020;7(6):1012–1023. Available: 10.1093/ 21(1):119–129.
nsr/nwaa036; https://doi.org/10.1093/nsr/nwaa036 Available: 10.1080/14737159.2021.1865807;
5. Andersen KG, Rambaut A, Lipkin WI, Holmes EC, https://doi.org/10.1080/14737159.2021.1865807
Garry RF. The proximal origin of SARS-CoV-2. 15. Nyaruaba R, Li X, Mwaliko C, et al. Two-step
Nat Med. 2020;26(4):450–452. reverse transcription droplet digital PCR protocols
Available: 10.1038/s41591-020-0820-9; for SARS-CoV-2 detection and quantification.
https://doi.org/10.1038/s41591-020-0820-9 JoVE. 2021;(169). Available: 10.3791/62295;
6. Wang M, Cao R, Zhang L, et al. Remdesivir and https://doi.org/10.3791/62295
chloroquine effectively inhibit the recently 16. Corman VM, Landt O, Kaiser M, et al. Detection
emerged novel coronavirus (2019-nCoV) in vitro. of 2019 novel coronavirus (2019-nCoV) by real-
Cell Res. 2020;30(3):269–271. time RT-PCR. Euro Surveill. 2020;25(3): 2000045.
Available: 10.1038/s41422-020-0282-0; Available: 10.2807/1560-
https://doi.org/10.1038/s41422-020-0282-0 7917.ES.2020.25.3.2000045;
7. Coronaviridae Study Group of the International https://doi.org/10.2807/1560-
Committee on Taxonomy of Viruses. The species 7917.ES.2020.25.3.2000045
severe acute respiratory syndrome-related 17. Chu DKW, Pan Y, Cheng SMS, et al. Molecular
coronavirus: classifying 2019-nCoV and naming it diagnosis of a novel coronavirus (2019-nCoV)
SARS-CoV-2. Nat Microbiol. 2020;5(4): 536–544. causing an outbreak of pneumonia. Clin Chem.
Available: 10.1038/s41564-020-0695-z; 2020;66(4):549–555.
https://doi.org/10.1038/s41564-020-0695-z Available: 10.1093/clinchem/hvaa029;
8. Huang C, Wang Y, Li X, et al. Clinical features of https://doi.org/10.1093/clinchem/hvaa029
patients infected with 2019 novel coronavirus in 18. Vogels CBF, Brito AF, Wyllie AL, et al. Analytical
Wuhan, China. Lancet. 2020;395(10223):497–506. sensitivity and efficiency comparisons of SARS-
Available: 10.1016/S0140-6736(20)30183-5; CoV-2 RT-qPCR primer-probe sets. Nat Microbiol.
https://doi.org/10.1016/S0140-6736(20)30183-5 2020;5(10):1299–1305.
9. Shi H, Han X, Jiang N, et al. Radiological findings Available: 10.1038/s41564-020-0761-6;
from 81 patients with COVID-19 pneumonia in https://doi.org/10.1038/s41564-020-0761-6
Wuhan, China: a descriptive study. Lancet Infect 19. Kralik P, Ricchi M. A basic guide to real time PCR
Dis. 2020;20(4):425–434. in microbial diagnostics: definitions, parameters,
Available: 10.1016/S1473-3099(20)30086-4; and everything. Front Microbiol. 2017;8:108.
https://doi.org/10.1016/S1473-3099(20)30086-4 Available: 10.3389/fmicb.2017.00108;
10. World Health Organization. Coronavirus disease https://doi.org/10.3389/fmicb.2017.00108
(COVID-19) weekly epidemiological update and 20. Fan Y, Zhao K, Shi ZL, Zhou P. Bat coronaviruses
weekly operational update. in China. Viruses. 2019;11(3):210.
https://www.who.int/emergencies/diseases/novel- Available: 10.3390/v11030210;
coronavirus-2019/situation-reports/ https://doi.org/10.3390/v11030210
11. Sun K, Chen J, Viboud C. Early epidemiological 21. Cheng VCC, Lau SKP, Woo PCY, Yuen KY. Severe
analysis of the coronavirus disease 2019 outbreak acute respiratory syndrome coronavirus as an agent
based on crowdsourced data: a population-level of emerging and reemerging infection. Clin
observational study. Lancet Digit Heal. Microbiol Rev 2007;20(4):660–694.
2020 ;2(4):e201–e208. Available: 10.1128/CMR.00023-07;
Available: 10.1016/S2589-7500(20)30026-1; https://doi.org/10.1128/CMR.00023-07
https://doi.org/10.1016/S2589-7500(20)30026-1 22. Chan JFW, Choi GKY, Tsang AKL, et al.
12. Wang FS, Zhang C. What to do next to control the Development and evaluation of novel real-time
2019-nCoV epidemic? Lancet. 2020;395 reverse transcription-PCR assays with locked
(10222):391–393. nucleic acid probes targeting leader sequences of
Available: 10.1016/S0140-6736(20)30300-7; human-pathogenic coronaviruses. J Clin Microbiol.
https://doi.org/10.1016/S0140-6736(20)30300-7 2015;53(8):2722–2726.
13. Zhang Y, Odiwuor N, Xiong J, et al. Rapid Available: 10.1128/JCM.01224-15;
molecular detection of SARS-CoV-2 (COVID-19) https://doi.org/10.1128/JCM.01224-15
virus RNA using colorimetric LAMP. medRxiv. 23. Lu X, Whitaker B, Sakthivel SKK, et al. Real-time
Available: 10.1101/2020.02.26.20028373; reverse transcription-PCR assay panel for Middle
https://doi.org/10.1101/2020.02.26.20028373 East respiratory syndrome oronavirus. J Clin
Submit a manuscript: https://www.tmrjournals.com/lr 9
doi: 10.53388/life2021-0525-110 ARTICLE
Microbiol. 2014;52(1):67– 75.
Available: 10.1128/JCM.02533-13;
https://doi.org/10.1128/JCM.02533-13
24. Chen HS, Su MJ, Shyu FM, et al. Mobile hospital:
healthcare for anybody in anytime and anywhere.
Proc 7th Int Work Enterp Netw Comput Healthc
Ind. 2005;2005:144–149.
Available: 10.1109/HEALTH.2005.1500425;
https://doi.org/10.1109/HEALTH.2005.1500425
25. Carossino M, Li Y, Lee PYA, et al. Evaluation of a
field-deployable reverse transcription-insulated
isothermal PCR for rapid and sensitive on-site
detection of Zika virus. BMC Infect Dis.
2017;17(1):778.
Available: 10.1186/s12879-017-2852-4;
https://doi.org/10.1186/s12879-017-2852-4
26. Wilkes RP, Lee PYA, Tsai YL, et al. An insulated
isothermal PCR method on a field-deployable
device for rapid and sensitive detection of canine
parvovirus type 2 at points of need. J Virol Methods.
2015;220:35–38.
Available: 10.1016/j.jviromet.2015.04.007;
https://doi.org/https://doi.org/10.1016/j.jviromet.2
015.04.007
27. Go YY, Kim YS, Cheon S, et al. Evaluation and
clinical validation of two field-deployable reverse
transcription-insulated isothermal PCR assays for
the detection of the Middle East respiratory
syndrome-coronavirus. J Mol Diagnostics.
2017;19(6):817–827.
Available: 10.1016/j.jmoldx.2017.06.007;
https://doi.org/10.1016/j.jmoldx.2017.06.007
28. Tracking coronavirus: Map, data and timeline -
BNO News.
https://bnonews.com/index.php/2020/02/the-
latest-coronavirus-cases/
29. Dong E, Du H, Gardner L. An interactive web-
based dashboard to track COVID-19 in real time.
Lancet Infect Dis. 2020;20(5):533–534.
Available: 10.1016/S1473-3099(20)30120-1;
https://doi.org/10.1016/S1473-3099(20)30120-1
30. Evonik industries - specialty chemicals -
productfinder.
https://corporate.evonik.com/en/products/search-
products/pages/product-
details.aspx?productId=27374&letter=R
31. Zowawi HM, Alenazi TH, AlOmaim WS, et al.
Portable RT-PCR system: a rapid and scalable
diagnostic tool for COVID-19 testing. J Clin
Microbiol. 2021;59(5):e03004–20.
Available: 10.1128/JCM.03004-20;
https://doi.org/10.1128/JCM.03004-20
Reviewer information Life Research thanks Daniel
Handley, K. Rajeshwar Reddy and other anonymous
reviewers for their contribution to the peer review of
this paper.
10 Submit a manuscript: https://www.tmrjournals.com/lr
You might also like
- Health Education HTPDocument4 pagesHealth Education HTPizelelelsNo ratings yet
- Hematology MCQsDocument84 pagesHematology MCQsHyder Ali84% (19)
- Lithium Carbonate Drug StudyDocument3 pagesLithium Carbonate Drug Studynot your medz duranNo ratings yet
- L'Helgouach-2020-EasyCOV - LAMP Based Rapid deDocument14 pagesL'Helgouach-2020-EasyCOV - LAMP Based Rapid deGerman MadrigalNo ratings yet
- Journal Pre-Proof: Journal of Clinical VirologyDocument12 pagesJournal Pre-Proof: Journal of Clinical Virologyamer khanNo ratings yet
- Evaluation of An Antigen Detection Rapid Diagnostic Test For Detection of Sars-Cov-2 in Clinical SamplesDocument9 pagesEvaluation of An Antigen Detection Rapid Diagnostic Test For Detection of Sars-Cov-2 in Clinical Samplesjytj tjrj64fdNo ratings yet
- Panbio COVID-19 Ag Rapid Test For The Diagnosis of SARS-CoV-2 - ArticuloDocument6 pagesPanbio COVID-19 Ag Rapid Test For The Diagnosis of SARS-CoV-2 - Articulo0244761No ratings yet
- Sensitive On-Site Detection of SARS-CoV-2 by ID NOW COVID-19Document12 pagesSensitive On-Site Detection of SARS-CoV-2 by ID NOW COVID-19luis.achaNo ratings yet
- 2019 New Coronavirus Nucleic Acid Detection Laboratory Construction Plan - 20200323Document15 pages2019 New Coronavirus Nucleic Acid Detection Laboratory Construction Plan - 20200323IMUNISASI DINKESNo ratings yet
- MainDocument7 pagesMainJhessica Gonzales GarciaNo ratings yet
- Approach To Assessment of New Swabs and Viral Transport Media For SARS-CoV-2 Testing-JCM.01562-20Document7 pagesApproach To Assessment of New Swabs and Viral Transport Media For SARS-CoV-2 Testing-JCM.01562-20Medtech ChuongNo ratings yet
- Lmo011 04 290Document7 pagesLmo011 04 290aobeso002No ratings yet
- Covid19 Methods PDFDocument14 pagesCovid19 Methods PDFSulieman OmarNo ratings yet
- Jurnal MPIDocument4 pagesJurnal MPIMarini TaslimaNo ratings yet
- 10 1128@JCM 00512-20Document22 pages10 1128@JCM 00512-20Luis KrNo ratings yet
- Swab Pooling For Large-Scale RT-QPCR Screening of Sars-Cov-2Document22 pagesSwab Pooling For Large-Scale RT-QPCR Screening of Sars-Cov-2popov357No ratings yet
- Quality in Diagnostic MicrobiologyDocument6 pagesQuality in Diagnostic Microbiologymicrobehunter007No ratings yet
- Multicenter Evaluation of The Panbio™ COVID-19 Rapid Antigendetection Test For The Diagnosis of SARS-CoV-2 InfectionDocument5 pagesMulticenter Evaluation of The Panbio™ COVID-19 Rapid Antigendetection Test For The Diagnosis of SARS-CoV-2 InfectionamyNo ratings yet
- JurnalDocument6 pagesJurnalDaffa AthaNo ratings yet
- Cid RT PCR Positive Over TimeDocument12 pagesCid RT PCR Positive Over TimePatrick CoNo ratings yet
- COVID-19 Testing One Size Does Not Fit AllDocument3 pagesCOVID-19 Testing One Size Does Not Fit Alliq_dianaNo ratings yet
- Molecular and Antibody Point-Of-Care Tests To Support The Screening, Diagnosis and Monitoring of COVID-19Document12 pagesMolecular and Antibody Point-Of-Care Tests To Support The Screening, Diagnosis and Monitoring of COVID-19vicndubNo ratings yet
- Biosafety Guidelines For The Use of GeneXperts As A Point-Of-Care Test (POCT)Document4 pagesBiosafety Guidelines For The Use of GeneXperts As A Point-Of-Care Test (POCT)katrinatigaNo ratings yet
- Pooling of Samples For Sars-Cov-2 Detection Using A Rapid Antigen TestDocument5 pagesPooling of Samples For Sars-Cov-2 Detection Using A Rapid Antigen TestErickson OngNo ratings yet
- The Detection Dogs Test Is More Sensitive Than Real-Time PCR in Screening For SARS-CoV-2Document7 pagesThe Detection Dogs Test Is More Sensitive Than Real-Time PCR in Screening For SARS-CoV-2Alejandro MerelesNo ratings yet
- Estudio Corona 1 Ropa41-7-E67Document5 pagesEstudio Corona 1 Ropa41-7-E67eudaldoNo ratings yet
- The Identification of The Sars Co V 2 Whole Genome Nine Cases Among Patients in Banten Province IndonesiaDocument13 pagesThe Identification of The Sars Co V 2 Whole Genome Nine Cases Among Patients in Banten Province IndonesiaTyas AnastasyaNo ratings yet
- Challenges in Laboratory Diagnosis of The NovelDocument27 pagesChallenges in Laboratory Diagnosis of The NovelRidho Al Fiqri100% (1)
- Title: Antigen Rapid Tests, Nasopharyngeal PCR and Saliva PCR To Detect Sars-Cov-2: A Prospective Comparative Clinical TrialDocument16 pagesTitle: Antigen Rapid Tests, Nasopharyngeal PCR and Saliva PCR To Detect Sars-Cov-2: A Prospective Comparative Clinical Trialorkut verificationNo ratings yet
- AbbottDocument10 pagesAbbottSisca PrimaNo ratings yet
- 108820lateral Move Molecular Assay For Horse StranglesDocument3 pages108820lateral Move Molecular Assay For Horse StranglesneriktjpcoNo ratings yet
- 1 s2.0 S0033350620301141 MainDocument3 pages1 s2.0 S0033350620301141 MainFedrik Monte Kristo LimbongNo ratings yet
- Real-Time RT-PCR For COVID-19 Diagnosis: Challenges and ProspectsDocument3 pagesReal-Time RT-PCR For COVID-19 Diagnosis: Challenges and ProspectsAnthony StackNo ratings yet
- CT Is Not EnoughDocument4 pagesCT Is Not Enougharhur9719No ratings yet
- Supardi, 2021Document15 pagesSupardi, 2021Khumaira SantaNo ratings yet
- Understanding Antigen Tests and Results ENG FinalDocument4 pagesUnderstanding Antigen Tests and Results ENG FinalAna CatarinaNo ratings yet
- JCM 01713-20Document13 pagesJCM 01713-20Yogiswara KarangNo ratings yet
- Genes 11 00664 v2Document13 pagesGenes 11 00664 v2Monika GonzalezNo ratings yet
- Assessing A Chip Based Rapid RTPCR Test For Sars Cov-2 Detection (Truenat Assay) : A Diagnostic Accuracy StudyDocument6 pagesAssessing A Chip Based Rapid RTPCR Test For Sars Cov-2 Detection (Truenat Assay) : A Diagnostic Accuracy StudySumesh Shreekhanda ShresthaNo ratings yet
- Serologic Responses To Sars-Cov-2 Infection Among Hospital Staff With Mild Disease in Eastern FranceDocument21 pagesSerologic Responses To Sars-Cov-2 Infection Among Hospital Staff With Mild Disease in Eastern FranceAssia AouachriaNo ratings yet
- Development of A Novel Loop-Mediated Isothermal Amplification Method For The Rapid Detection of Monkeypox Virus InfectionsDocument12 pagesDevelopment of A Novel Loop-Mediated Isothermal Amplification Method For The Rapid Detection of Monkeypox Virus InfectionsV K kasaragodNo ratings yet
- Predicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 From Diagnostic SamplesDocument4 pagesPredicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 From Diagnostic SamplesmeNo ratings yet
- Up Date Diagnosis Laboratorium Pada Pedoman: Pencegahan Dan Pengendalian COVID-19 Revisi Ke-5Document65 pagesUp Date Diagnosis Laboratorium Pada Pedoman: Pencegahan Dan Pengendalian COVID-19 Revisi Ke-5Muhammad NurfiantoroNo ratings yet
- Predicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 From Diagnostic SamplesDocument4 pagesPredicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 From Diagnostic SamplesArum Rahmawatie VirgieNo ratings yet
- Jurnal 2Document10 pagesJurnal 2GFORCE .ID.No ratings yet
- ! Rapid Detection of Novel Coronavirus (COVID-19) by Reverse Transcription-Loop-Mediated Isothermal AmplificationDocument17 pages! Rapid Detection of Novel Coronavirus (COVID-19) by Reverse Transcription-Loop-Mediated Isothermal AmplificationPetr CiglerNo ratings yet
- Presence of Mismatches Between Diagnostic PCR Assays and Coronavirus Sars-Cov-2 GenomeDocument17 pagesPresence of Mismatches Between Diagnostic PCR Assays and Coronavirus Sars-Cov-2 GenomeYulaimy BatistaNo ratings yet
- Journal of Clinical Microbiology-2020-Chan-JCM.00310-20.full PDFDocument33 pagesJournal of Clinical Microbiology-2020-Chan-JCM.00310-20.full PDFDwi Ratna KarimNo ratings yet
- Multiplex Detection and Dynamics of IgG Antibodies To SARS-CoV2 and The Highly Pathogenic Human Coronaviruses SARS-CoV and MERS-CoVDocument7 pagesMultiplex Detection and Dynamics of IgG Antibodies To SARS-CoV2 and The Highly Pathogenic Human Coronaviruses SARS-CoV and MERS-CoVamyNo ratings yet
- 2020 Article 335 PDFDocument4 pages2020 Article 335 PDFFellita Ratri ANo ratings yet
- Makalah Biologi MolekulerDocument11 pagesMakalah Biologi MolekulerSarah PanjaitanNo ratings yet
- OASH COVID 19 Guidance Testing PlatformsDocument3 pagesOASH COVID 19 Guidance Testing PlatformsNurul PalesseiNo ratings yet
- Journal of Clinical Microbiology-2020-Chan-JCM.00310-20.full PDFDocument33 pagesJournal of Clinical Microbiology-2020-Chan-JCM.00310-20.full PDFdavid zhangNo ratings yet
- Ijerph-18-11451 Meta Analysis StudyDocument14 pagesIjerph-18-11451 Meta Analysis StudySumit BediNo ratings yet
- Different Longitudinal Patterns of Nucleic Acid and Serology Testing Results Based On Disease Severity of COVID-19 PatientsDocument5 pagesDifferent Longitudinal Patterns of Nucleic Acid and Serology Testing Results Based On Disease Severity of COVID-19 PatientsagboxNo ratings yet
- Towards Detection of SARS-CoV-2 RNADocument8 pagesTowards Detection of SARS-CoV-2 RNARuben MonroyNo ratings yet
- Correspondence To: Professor Changxin Shen Department of Blood TransfusionDocument13 pagesCorrespondence To: Professor Changxin Shen Department of Blood TransfusionAlfa RidziNo ratings yet
- Lynx PublicationDocument9 pagesLynx PublicationSwarnendraNo ratings yet
- Performance Verification of Anti-Sars-Cov-2-Specific Antibody Detection by Using Four Chemiluminescence Immunoassay SystemsDocument6 pagesPerformance Verification of Anti-Sars-Cov-2-Specific Antibody Detection by Using Four Chemiluminescence Immunoassay SystemsadnanNo ratings yet
- A Role For Biofoundries in Rapid Development and Validation of Automated Sars-Cov-2 Clinical DiagnosticsDocument11 pagesA Role For Biofoundries in Rapid Development and Validation of Automated Sars-Cov-2 Clinical DiagnosticsRicardoPeixotoNo ratings yet
- Validasi Rapid AntigenDocument6 pagesValidasi Rapid Antigengaki telitiNo ratings yet
- Artigo Cientifico Barreto2020 Article Diagnosingthenovelsars Cov 2bypdfDocument10 pagesArtigo Cientifico Barreto2020 Article Diagnosingthenovelsars Cov 2bypdfOtaku GamerChanNo ratings yet
- SARS-CoV-2 Viral Outbreak Investigation: Laboratory Perspective: Clinical Updates in COVID-19From EverandSARS-CoV-2 Viral Outbreak Investigation: Laboratory Perspective: Clinical Updates in COVID-19Rating: 3 out of 5 stars3/5 (1)
- Athlete's Heart: Dr. Arzalan BaigDocument59 pagesAthlete's Heart: Dr. Arzalan BaigArzalan BaigNo ratings yet
- TRẮC NGHIỆM AVCN PHARMACYDocument91 pagesTRẮC NGHIỆM AVCN PHARMACYHoang Pham ThaiNo ratings yet
- Fisiologia Do EnvelhecimentoDocument2 pagesFisiologia Do EnvelhecimentoLissandro DannyNo ratings yet
- Physical Examination FormatDocument6 pagesPhysical Examination Formatvethadhas vin100% (1)
- Student'S Name: Kim Christian P. Pagobo Subject: Science 120: Anatomy and Physciology Instructor: Mrs. Roma Grace DizonDocument9 pagesStudent'S Name: Kim Christian P. Pagobo Subject: Science 120: Anatomy and Physciology Instructor: Mrs. Roma Grace Dizonkim christianNo ratings yet
- Coronary Artery Diseases - Ilham PDocument43 pagesCoronary Artery Diseases - Ilham PKeputrian FKUPNo ratings yet
- Patient Classification & AssignmentDocument37 pagesPatient Classification & AssignmentAkhila UsNo ratings yet
- J.H. Cerilles State College: Western Mindanao State UniversityDocument6 pagesJ.H. Cerilles State College: Western Mindanao State UniversityVanessa PalomaNo ratings yet
- Nclex Study GuideDocument37 pagesNclex Study GuideBlas Christian Tancongco ChavesNo ratings yet
- Fermented Products With Probiotic Qualities - Casei & Bifidobakterije PDFDocument6 pagesFermented Products With Probiotic Qualities - Casei & Bifidobakterije PDFmilu1312No ratings yet
- Bigay-Panghulan - Legal Opinion - Legal Forms - First ActivityDocument3 pagesBigay-Panghulan - Legal Opinion - Legal Forms - First ActivityApril Ann Sapinoso Bigay-PanghulanNo ratings yet
- UHSAA Preparticipation FormDocument4 pagesUHSAA Preparticipation FormThe Salt Lake TribuneNo ratings yet
- Functional Constipation: A Case Report: EpartmentDocument5 pagesFunctional Constipation: A Case Report: EpartmentArielIzaSuquilloNo ratings yet
- Univestin+ Glucosmine NewDocument26 pagesUnivestin+ Glucosmine NewAshishNo ratings yet
- Finalised ILERA EKO DIASPORA PLAN PRODUCT PAPER 1Document19 pagesFinalised ILERA EKO DIASPORA PLAN PRODUCT PAPER 1dibransonsNo ratings yet
- Donning Doffing Spotter Check List v1Document1 pageDonning Doffing Spotter Check List v1Rudyard Paul AmistadNo ratings yet
- Lab 2,3Document18 pagesLab 2,3Nosheela KhalidNo ratings yet
- New Patient Nutrition Assessment Form: (Person 1)Document12 pagesNew Patient Nutrition Assessment Form: (Person 1)Ayesha KhanNo ratings yet
- Krauses Food and The Nutrition Care Process 14Th Edition Mahan Test Bank Full Chapter PDFDocument28 pagesKrauses Food and The Nutrition Care Process 14Th Edition Mahan Test Bank Full Chapter PDFphualexandrad39100% (12)
- Blaylock Hypertension 13Document16 pagesBlaylock Hypertension 13mohanp71No ratings yet
- Bloodbld 2019003808 CDocument12 pagesBloodbld 2019003808 CBillal BenhaddadNo ratings yet
- OSCES 5e MMSDocument125 pagesOSCES 5e MMSFlórian Jefferson100% (1)
- Vinblastine (Velban®) : Things That May Occur During or Within Hours of TreatmentDocument2 pagesVinblastine (Velban®) : Things That May Occur During or Within Hours of TreatmentVette Angelikka Dela CruzNo ratings yet
- Rehan Aftab Hamdulay, Himanshu Girish Sharma, Bombay College of Pharmacy, Santacruz (East), Mumbai - 400098Document1 pageRehan Aftab Hamdulay, Himanshu Girish Sharma, Bombay College of Pharmacy, Santacruz (East), Mumbai - 400098Himanshu SharmaNo ratings yet
- Inguinal Hernia Case FileDocument2 pagesInguinal Hernia Case Filehttps://medical-phd.blogspot.comNo ratings yet
- Pranav SharmaDocument11 pagesPranav SharmaPranav SharmaNo ratings yet
- Kapalbhati PDFDocument2 pagesKapalbhati PDFRahul MishraNo ratings yet