Professional Documents
Culture Documents
Application of Isothermal Titration Calorimetry in The Biological Sciences: Things Are Heating Up!
Application of Isothermal Titration Calorimetry in The Biological Sciences: Things Are Heating Up!
Uploaded by
Jack HsuOriginal Description:
Original Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Application of Isothermal Titration Calorimetry in The Biological Sciences: Things Are Heating Up!
Application of Isothermal Titration Calorimetry in The Biological Sciences: Things Are Heating Up!
Uploaded by
Jack HsuCopyright:
Available Formats
Application of Isothermal Titration Calorimetry in the Biological Sciences: Things Are Heating Up!
John E. Ladbury University College London, London, UK
During the last 15 years isothermal titration calorimetry (ITC) has come of age. There are now in excess of 2000 instruments sited in laboratories in more than 40 countries around the world. Research scientists in such diverse elds as biophysics, cell biology, pharmaceutical screening, and food research routinely investigate their systems of interest using ITC. Why is it that this methodology has sparked such enthusiasm and interest, and what use is the data obtained? The dramatic advances in the eld of structural biology in the last couple of decades fed a desire of biochemists to dene molecular function and mechanism in ever increasing detail. Describing a biochemical process however, cannot be served by structure alone. A full understanding is only obtained with a quantication of the change of state of the system. In an equilibrium process, such as a biomolecular interaction, thermodynamic measurement provides quantication of the change in energy on going from the free to the bound state. The ITC instrument (for reviews, see References 15) uses the extremely accurate measurement of heat as a probe for an interaction as it occurs. Knowing the concentrations of the interacting moieties allows the calculation of the observed change in molar enthalpy of the interaction, Hobs. The term observed (denoted by the subscript obs) signies that the quantity is not solely from the isolated events associated with forming a biomolecular interface (i.e., direct noncovalent bonds between the atoms of the interacting moieties), but also includes heat derived from perturbation of the solvent around the binding site, potential conformational changes occurring elsewhere in the interacting biomolecules (6), and direct formation of noncovalent bonds between other solutes such as ions or apolar compounds that may be incorporated as an ingredient of the bulk solvent. Since every biomolecular interaction has either an uptake or release of heat associated with it, the ITC is a universal detector of the occurrence of binding (at an appropriate temperature). The direct determination of the Hobs negates the indirect calculation of this parameter using a vant Hoff-based method, which can be problematic over extended temperature ranges due to the inuence of the change in heat capacity (i.e., the H changes with temperature; see Equation 3). Furthermore, since the two components of an interaction can be titrated, the measured heat gives a direct readout of the extent of interaction at any given concentration regime (see Figure 1 and Reference 7 for experimental tutorial). As a result, the concentrations of free and bound molecules and hence the observed binding or dissociation constant, (KB,obs or KD,obs, respectively; KB = 1/KD) can be determined. Armed with the Hobs and the KB,obs, a full thermodynamic description of the interaction can be elucidated at a given experimental temperature (T) based on the following relationships: Gobs = -RTln KB,obs [Eq. 1]
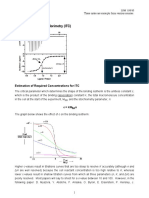
Figure 1. Schematic representations of isothermal titration calorimetry (ITC) instruments. (A) An ITC instrument prior to performing a titration. The sample cell and the reference cell as kept at the same temperature, which is typically 510C above the temperature maintained outside the jacket in which the cells are housed. The reference cell is always kept at the experimental temperature. One of the components of the interaction is placed in the syringe and the other in the cell. (B) A ITC instrument performing a titration. When an injection is made, the change in heat associated with binding (endothermic or exothermic) results in a change in temperature in the sample cell. A change in power (heat/s) is required to return the cells to identical temperatures (T) (i.e., T = 0). This change in power is recorded as a series of injections is made. In the raw data presented in the inset, each injection is accompanied by an interaction where heat is given out (exothermic). As the course of injections is completed, the binding sites on the sample in the cell are gradually saturated, and the exothermic effect becomes reduced. For details, see References 15 and 7. Vol. 37, No. 6 (2004) BioTechniques 885
Table 1. ITC Measurement of the Interaction of the Src SH2 Domain With a Series of Peptides Peptide Sequence Ile Val Glu Trp Asp KB 106 M-1 10.80 6.24 4.87 3.18 2.64 G (kJ/mol) -40.1 -38.8 -38.1 -37.1 -36.6 H (kJ/mol) -38.7 -28.6 -32.7 -32.2 -27.5 TS (kJ/mol) 1.4 10.2 5.4 4.9 9.1
a range of pH. The dependence of the KB,obs on the pH can reveal what protonation events might be occurring on binding and their respective pKa values (10,12). Furthermore, the relationship between the Hobs and the change in enthalpies for the experiment performed in dened buffer solutions (with differing heats of ionization) can indicate the exact number of protons in exchange (10). A similar understanding of the electrostatic contribution to the formation of a complex can be determined by performing the ITC experiment over a range of salt concentrations. The uptake, or release of ions, can be observed in the dependence of the KB,obs on the concentration of salt (see Figure 2 and Reference 13).
Data taken from Reference 11. All peptides have the sequence Ac-GluProGln(pTyr)GluGluXxxProIleTyrLeu-NH2, where Xxx is the residue denoted in the first column of the Table. ITC, isothermal titration calorimetry; KB, observed binding constant; G, change in Gibbs free energy; H, change in observed enthalpy; T, experimental temperature; S, change in entropy.
The Correlation of Thermodynamics with Biomolecular Structure
Based on an empirically determined correlation of the Cp term with the change in surface area buried on forming a bimolecular interface, thermodynamic parameters can be directly related to high resolution structure, and prediction of one from the other is possible (14,15). ITC experiments performed at different temperatures provide an accurate direct determination of the Cp term (see Equation 3). It should be pointed out that this correlation should come with a warning that it can be misleading in cases, for example, where water molecules appear in the interface (13,16,17) or where signicant conformational changes in one or both molecules accompany the binding (18). Another important advantage of the full thermodynamic characterization provided by ITC is the additional level of information by which to distinguish binding events. In some instances,
from which the change in Gibbs free energy (Gobs) is obtained (where R is the gas constant), and Sobs = (Hobs - Gobs)/T [Eq. 2]
from which the change in entropy (Sobs) can be ascertained. A further thermodynamic characterization of an interaction, the change in constant pressure heat capacity (Cp) can be determined from measurement of the H over a range of temperatures since Cp = dHobs /dT [Eq. 3]
Thus ITC has become the instrument of choice for thermodynamic analysis of biomolecular interactions, since it can provide the H, G, and the S in one experiment. Furthermore, the experiment is rapid, typically taking less than an hour, there is no need for chemical modication or immobilization of any of the components, there is no size limitation on the molecules to be investigated, and the experiment can be conducted in turbid or colored solutions or in the presence of suspensions.
Why Use ITC?
The complete thermodynamic characterization of a bimolecular interaction can be obtained relatively easily using the ITC in one experiment. However, the assimilation of these data is not so straightforward. The underlying events contributing to a biological interaction are many fold and interlinked in a complicated manner. The observed thermodynamic parameters associated with forming a series of noncovalent bonds between the biomolecules in forming a complex cannot be divorced from those derived from breaking bonds as the solvent rearranges to accommodate the complex. The latter effect cannot currently be determined empirically. Therefore, it is reasonable to question how data derived from ITC can add to our understanding of biology. The answer to this question lies in the ability to separate out various contributions to binding using a combination of ITC experiments. The following serves exemplify this approach.
pH and Electrostatic Effects on Complex Formation
A single ITC experiment does not provide thermodynamic data for an isolated bond (or series of bonds). However, there are methods to deconvolve (or parse) various contributions occurring on formation of a complex such that a detailed picture of the energetics that drive the interaction can be determined (811). For example, the thermodynamic parameters associated with a protonation/deprotonation event(s) on forming a biomolecular complex can be determined by performing titrations over
886 BioTechniques
Figure 2. Log KB, obs against the log of the salt concentration for the interaction of the archaeal Pyrococcus woesei TATA binding protein (PwTBP) with DNA. The dependence of the equilibrium binding constant, KB,obs, on the concentration of a monovalent salt, [MX], can be determined using the relationship: logKB,obs = logKref - Alog[MX] + 0.16B[MX] where A is the net number of ions released (a negative number) or taken up (a positive number) on forming the complex, B is the total number of water molecules released (a positive number) on forming the complex, and Kref is a hypothetical binding constant at 1 M salt, in the absence of any contribution from release or uptake of water (i.e., B = 0). Binding studies on the PwTBP DNA interaction, performed at high salt concentrations using isothermal titration calorimetry (ITC), reveal that the interaction occurs with a net uptake of two ions and the release of approximately 40 water molecules (for details, see Reference 13). Vol. 37, No. 6 (2004)
the KB of a series of interactions with one biomolecule are similar or indistinguishable, however, the determination of the H and S terms allows a further level of discrimination. Although this may not provide a rigorous denition of the changes in noncovalent interactions on the atomic level, it can nonetheless be used as a guideline in understanding binding. For example, the interaction of a series of ligands with the SH2 domain from the protein Src show very similar afnities (between 2.6410.8 M-1), however the enthalpic and entropic changes that occur provide a clear level of discrimination between the interactions (see Table 1 and Reference 19). These data reveal that the tightest interaction has a signicantly lower favorable (+ve) TS term and a more favorable (-ve) H term. Whereas substituting the Ile residue for a Val dramatically improves the TS value, but the compensating H is signicantly less favorable. Based on the high resolution structural detail, it is possible to speculate that the placing of the smaller Val side chain in a pocket that accommodates the Ile results in fewer noncovalent bonds (less favorable H, but allows the Val to occupy more congurational states; favorable S). The subtle difference between these two peptides has a limited effect on the afnity but clearly has a dramatic effect on the way in which binding occurs, which is revealed by the H and TS terms. Thus, although, it may not be possible to provide a full thermodynamic-structure correlation from an ITC experiment, it is possible to draw sensible conclusions from data by comparing systems where subtle changes have been imposed.
References
1. Wiseman, T., S. Williston, J.F. Brandts, and L.N. Lin. 1989. Rapid measurement of binding constants and heats of binding using a new titration calorimeter. Anal. Biochem. 179:131-137. 2. Freire, E., O.L. Mayorga, and M. Straume. 1990. Isothermal titration calorimetry. Anal. Chem. 62:950-959. 3. Ladbury, J.E. 1995. Counting the calories to stay in the groove. Structure 3:635-639. 4. Ladbury, J.E. and B.Z. Chowdhry. 1996. Sensing the heat: the application of isothermal titration calorimetry to thermodynamic studies of biomolecular interactions. Chem. Biol. 3:791-801. 5. Jelasarov, I. and H.R. Bosshard. 1999. Isothermal titration calorimetry and differential scanning calorimetry as complementary tools to investigate the energetics of biomolecular recognition. J. Mol. Recogn. 12:3-18. 6. Williams, M.A. and J.E. Ladbury. 2004. The extended interface: measuring non-local effects in biomolecular interactions. Curr. Opin Struct. Biol. 14:562-569. 7. Thomson, J.A. and J.E. Ladbury. 2004. Isothermal titration calorimetry: a tutorial. In J.E. Ladbury and M.L. Doyle (Eds.), Biocalorimetry 2. Applications of Calorimetry in the Biological Sciences. John Wiley & Sons, Chichester. 8. Haq, I., J.E. Ladbury, B.Z. Chowdhry, T.C. Jenkins, and J.B. Chaires. 1997. Specific binding of Hoechst 33258 to the d(GGCAAATTTGCG)2 duplex: calorimetric and spectroscopic studies. J. Mol. Biol. 271:244-257. 9. Luque, I., M.J. Todd, J. Gomez, N. Semo, and E. Freire. 1998. Molecular basis of resistance to HIV-1 protease inhibition: a plausible hypothesis. Biochemistry 37:5791-5797. 10. Baker, B.M. and K.P. Murphy. 1996. Evaluation of linked protonation effects in protein binding reactions using isothermal titration calorimetry. Biophys. J. 71:2049-2055. 11. Ren, J.S., T.C. Jenkins, and J.B. Chaires. 2000. Energetics of DNA intercalation reactions. Biochemistry 39:8439-8447. 12. Haq, I., R. OBrien, A. Lagunavicius, V. Siksnys, and J. Ladbury. 2001. Specific DNA recognition by the type II restriction endonuclease MunI: the effect of pH. Biochemistry 40:14960-14967. 13. Bergqvist, S., M.A. Williams, R. OBrien, and J.E. Ladbury. 2004. Heat capacity effects of water molecules and ions at a protein-DNA interface. J. Mol. Biol. 336:829-842. 14. Spolar, R.S. and M.T. Record, Jr. 1994. Coupling of local folding to sitespecific binding of proteins to DNA. Science 263:777-783. 15. Gomez, J. and E. Freire. 1995 Thermodynamic mapping of the inhibitor site of the aspartic protease endothiapepsin. J. Mol. Biol. 252:337-350. 16. Morton, C.J. and J.E. Ladbury. 1996. Water mediated protein-DNA interactions: the relationship of thermodynamics to structural detail. Prot. Sci. 5:2115-2118. 17. Henriques, D.A., J.E. Ladbury, and R.M. Jackson. 2000. Comparison of binding energies of SrcSH2-phosphotyrosyl peptides with structurebased prediction using surface area based empirical parameterization. Prot. Sci. 9:1975-1985. 18. Thomson, J., G.S. Ratnaparkhi, R. Varadarajan, J.M. Sturtevant, and F.M. Richards. 1994. Thermodynamic and structural consequences of changing a sulphur atom to a methylene group in the M13Nle mutation in ribonuclease S. Biochemistry 33:8587-8593. 19. Henriques, D.A. and J.E. Ladbury. 2001. Inhibitors to the Src SH2 domain: a lesson in thermodynamic structural correlation in drug design. Arch. Biochem. Biophys. 390:158-168.
The Future of ITC
Technical advances in ITC instrumentation are likely to increase the viability of the technique in more research laboratories. Recent developments have largely centered on attempting to improve the throughput of samples such that ITC can be, at least partially, integrated into screening or drug development scenarios. Instrumentation that includes automated sample handling capability is currently available, and instruments with silicon chip-based heat sensors capable of analysis of small samples in a 96-well plate format are in development. The heat of interaction provides a signal that a desired molecule or hit from a screen has bound. The real strength of ITC in the development of pharmaceuticals is to add a further level of information to the decision-making process. For example, in drug development, it commonly occurs that decisions have to be made as to which compounds to pursue as leads from a large number of hit compounds derived from initial screens of libraries. In a case where two or more compounds have similar afnities, it has been suggested that the compounds with the most favorable H terms are likely to make the best lead compounds for further chemical modication. This is because the H term corresponds to the energy associated with the net change in noncovalent bonds. A more favorable H term suggests a better complementarity of bonds in the interface. Since improving the bonding in the biomolecule-drug interface is a major challenge, the initial enthalpy data is a useful tool to discriminate between compounds. Upping the ante in the race to develop new drug compounds requires as much help in deciding which molecules to pursue as possible. As a result, the addition of thermodynamic data in the decision making process will become widely adopted. It is unlikely that ITC will in the foreseeable future be able to make a valuable contribution to the identication of hits in highthroughput screens due to instrumental limitations and possible requirements of materials. However, the integration of thermodynamic data at the decision making point in secondary screening and in optimization of rational design will be fundamental applications.
Vol. 37, No. 6 (2004)
BioTechniques 887
You might also like
- Adiabatic Reactors Final Lab Group 1-ADocument22 pagesAdiabatic Reactors Final Lab Group 1-AHaris SheikhNo ratings yet
- Abstract For CSTR Lab ReportDocument4 pagesAbstract For CSTR Lab ReportNabilah SyaheeraNo ratings yet
- Isothermal Titration CalorimetryDocument6 pagesIsothermal Titration CalorimetryMiles NsgNo ratings yet
- Solvent Effects On The Singlet - Triplet Equilibrium and Reactivity of A Ground Triplet State Arylalkyl CarbeneDocument4 pagesSolvent Effects On The Singlet - Triplet Equilibrium and Reactivity of A Ground Triplet State Arylalkyl CarbeneSergioSilvaNo ratings yet
- Assistant Professor of Chemistry, B.S. College Hatta Chenari, V.K.S. University, AraDocument6 pagesAssistant Professor of Chemistry, B.S. College Hatta Chenari, V.K.S. University, AraAnonymous CwJeBCAXpNo ratings yet
- Itc Current ProtocolsDocument24 pagesItc Current ProtocolsRigel_TNo ratings yet
- (1990, Freire Et Al.) Isothermal Titration CalorimetryDocument10 pages(1990, Freire Et Al.) Isothermal Titration CalorimetryAlan R. F. K. MoraesNo ratings yet
- J. Patrick A. Fairclough Et Al - Chain Length Dependence of The Mean Ðeld Temperature in Poly (Oxyethylene) - Poly (Oxybutylene) Diblock CopolymersDocument3 pagesJ. Patrick A. Fairclough Et Al - Chain Length Dependence of The Mean Ðeld Temperature in Poly (Oxyethylene) - Poly (Oxybutylene) Diblock CopolymersDremHpNo ratings yet
- Paper Isomerization Nitrito Complejos CoDocument7 pagesPaper Isomerization Nitrito Complejos CoJuan Gabriel FernándezNo ratings yet
- 1982 - Mcelhaney - The Use of Differential Scanning Calorimetry and Differential Thermal Analysis in Studies of Model and Biological MembranesDocument31 pages1982 - Mcelhaney - The Use of Differential Scanning Calorimetry and Differential Thermal Analysis in Studies of Model and Biological MembranesymiyazyNo ratings yet
- Pharmaceutical Kinetics and Stability-Lec-5Document23 pagesPharmaceutical Kinetics and Stability-Lec-5husseinkamilhamid123456789No ratings yet
- Cool FlamesDocument12 pagesCool FlamesQasim IsmailNo ratings yet
- Equilibrium Constants Methyl Tert-Butyl Ether Liquid-Phase SynthesisDocument5 pagesEquilibrium Constants Methyl Tert-Butyl Ether Liquid-Phase Synthesisjulior87No ratings yet
- A of Of: Thermody:namic Study The Interaction Tubulin With Colchicine Ligands'@Document5 pagesA of Of: Thermody:namic Study The Interaction Tubulin With Colchicine Ligands'@Carlos LimacoNo ratings yet
- The Effect of Temperature On Enzyme Activity New IDocument10 pagesThe Effect of Temperature On Enzyme Activity New Ishikai towNo ratings yet
- Tesmar2016 Article BufferContributionToFormationEDocument6 pagesTesmar2016 Article BufferContributionToFormationEMuhammad AriviansyahNo ratings yet
- Screenshot 2022-10-16 at 7.16.17 PMDocument49 pagesScreenshot 2022-10-16 at 7.16.17 PMDanaNo ratings yet
- Bond Dissociation Energies For Diatomic Molecules Containing 3d Transition Metals: Benchmark Scalar Relativistic Coupled Cluster Calculations For Twenty MoleculesDocument40 pagesBond Dissociation Energies For Diatomic Molecules Containing 3d Transition Metals: Benchmark Scalar Relativistic Coupled Cluster Calculations For Twenty MoleculesNyau NyauNo ratings yet
- Interphase Mass Transfer: Demonstrating The Effect of in A Transparent Fluidized Bed Reactor Science - Gov (United States)Document6 pagesInterphase Mass Transfer: Demonstrating The Effect of in A Transparent Fluidized Bed Reactor Science - Gov (United States)Sandeepkumar SharmaNo ratings yet
- The Third Law of ThermodynamicsDocument10 pagesThe Third Law of ThermodynamicssamygoldNo ratings yet
- Energy Dissipation in Slipping Biologica PDFDocument10 pagesEnergy Dissipation in Slipping Biologica PDFVivi YantimalaNo ratings yet
- Voet 01Document11 pagesVoet 01Xilo0000No ratings yet
- Isothermal Titration Calorimetry: Presented By: Ms. Prajakta S.Pawar. Guided byDocument54 pagesIsothermal Titration Calorimetry: Presented By: Ms. Prajakta S.Pawar. Guided byVrushali Puranik-GokhaleNo ratings yet
- Screenshot 2022-10-09 at 6.48.23 PMDocument51 pagesScreenshot 2022-10-09 at 6.48.23 PMDanaNo ratings yet
- Corresponding State TheoryDocument15 pagesCorresponding State TheoryAravind KNo ratings yet
- Biopolymers - November 1982 - Marky - Calorimetric Determination of Base Stacking Enthalpies in Double Helical DNADocument10 pagesBiopolymers - November 1982 - Marky - Calorimetric Determination of Base Stacking Enthalpies in Double Helical DNAhajardreamhighNo ratings yet
- VLE Lactic Acid Ethyl Lactate Esterification PDFDocument7 pagesVLE Lactic Acid Ethyl Lactate Esterification PDFAseem Kashyap0% (1)
- 2009 WunderlichHeimburgSchneider BJDocument6 pages2009 WunderlichHeimburgSchneider BJKristy IngramNo ratings yet
- Energetics NoteDocument9 pagesEnergetics Notecupcakesen22No ratings yet
- Chem 16Document8 pagesChem 16Adi SoNo ratings yet
- Effect of Temperature On The Reaction RateDocument5 pagesEffect of Temperature On The Reaction RateChristy Joy RetanalNo ratings yet
- The Law of Entropy Invcrease - A Lab ExperimentDocument4 pagesThe Law of Entropy Invcrease - A Lab ExperimentINDUSTRIA BIOTECNOLÓGICA DEL SURNo ratings yet
- Electrodilatometry of Liquids, Binary Liquids, and SurfactantsDocument12 pagesElectrodilatometry of Liquids, Binary Liquids, and SurfactantsmilousNo ratings yet
- Supercritical Water Gasification of Biomass Thermodynamic AnalysisDocument7 pagesSupercritical Water Gasification of Biomass Thermodynamic AnalysisLuiz Guilherme SilvaNo ratings yet
- Thermodynamic Calculations For Biochemical Transport and Reaction Processes in Metabolic NetworksDocument9 pagesThermodynamic Calculations For Biochemical Transport and Reaction Processes in Metabolic NetworksDizaichaNo ratings yet
- Thermodynamic Calculations For Biochemical Transport and Reaction Processes in Metabolic NetworksDocument9 pagesThermodynamic Calculations For Biochemical Transport and Reaction Processes in Metabolic NetworksDizaichaNo ratings yet
- Un Nuevo Método para Explorar El Comportamiento Térmico y de Ventilación de La Fuga Térmica de La Batería de Iones de LitioDocument6 pagesUn Nuevo Método para Explorar El Comportamiento Térmico y de Ventilación de La Fuga Térmica de La Batería de Iones de LitioLaura Marcela Montoya GomezNo ratings yet
- Kumar Nussinov Biochem 2001 Mesophile Thermophile StabilityDocument14 pagesKumar Nussinov Biochem 2001 Mesophile Thermophile StabilityJames NanorNo ratings yet
- CO Oxidation On PT Variable Phasing of I PDFDocument11 pagesCO Oxidation On PT Variable Phasing of I PDFTysir SarhanNo ratings yet
- Differential Scanning Calorimetry in Food Research A ReviewDocument27 pagesDifferential Scanning Calorimetry in Food Research A ReviewMarius MihaiNo ratings yet
- Houk Hydroboration TetDocument18 pagesHouk Hydroboration TetSuavo Tekka MukherjeeNo ratings yet
- Thermodynamics of Microbial Growth and MetabolismDocument17 pagesThermodynamics of Microbial Growth and MetabolismJeimy MaciasNo ratings yet
- KOHDocument6 pagesKOHSoumen GuhaNo ratings yet
- Nahid Sohrevardi, Farhoush Kiani, Fardad KoohyarDocument12 pagesNahid Sohrevardi, Farhoush Kiani, Fardad KoohyarLu Pham KhacNo ratings yet
- 2011 - MST Technology ReviewDocument12 pages2011 - MST Technology Reviewhakos12No ratings yet
- A Thermodynamic Approach To The Problem of Stabilization of Globular Protein Structure A Calorimetric StudyDocument20 pagesA Thermodynamic Approach To The Problem of Stabilization of Globular Protein Structure A Calorimetric StudyCaroline LacerdaNo ratings yet
- 16 Distillation in The Pharmaceutical Industry: Rf. WilcoxDocument39 pages16 Distillation in The Pharmaceutical Industry: Rf. WilcoxcsandrasNo ratings yet
- Estequiometría Del Crecimiento Microbiano y Formación de ProductoDocument9 pagesEstequiometría Del Crecimiento Microbiano y Formación de ProductoDaniela NavasNo ratings yet
- 453 LiMin NgaifestDocument9 pages453 LiMin NgaifestNadie NingunoNo ratings yet
- Differential Scanning Calorimetry (DSC)Document9 pagesDifferential Scanning Calorimetry (DSC)DanielNo ratings yet
- FH y UNIQUACDocument28 pagesFH y UNIQUAClauraNo ratings yet
- Liquid Phase ReactorDocument22 pagesLiquid Phase Reactorkrishy19s100% (2)
- Nonclassical Chemical Kinetics For Description of Chemical Fluctuation in A Dynamically Heterogeneous Biological SystemDocument8 pagesNonclassical Chemical Kinetics For Description of Chemical Fluctuation in A Dynamically Heterogeneous Biological SystemIshwar ChandraNo ratings yet
- Formal Written Lab Report - Experiment 1 - Group 1 (1st Year - Summer Sem)Document7 pagesFormal Written Lab Report - Experiment 1 - Group 1 (1st Year - Summer Sem)Cheska BiolenaNo ratings yet
- Chemical KineticsDocument3 pagesChemical KineticsYEEHSHIN JILL GAYONo ratings yet
- Belfiore y Gomez-Garcia, Transport Phenomena For Chemical Rector Design - KOE 2018Document65 pagesBelfiore y Gomez-Garcia, Transport Phenomena For Chemical Rector Design - KOE 2018Zully CabreraNo ratings yet
- Designing Itc v7 ExptsDocument5 pagesDesigning Itc v7 ExptsRigel_TNo ratings yet
- Physico-Chemistry of Solid-Gas Interfaces: Concepts and Methodology for Gas Sensor DevelopmentFrom EverandPhysico-Chemistry of Solid-Gas Interfaces: Concepts and Methodology for Gas Sensor DevelopmentNo ratings yet
- QlikSense Course ContentDocument4 pagesQlikSense Course ContentSatyam GargNo ratings yet
- Abap On Cloud CNA385Document40 pagesAbap On Cloud CNA385shibuNo ratings yet
- Near Net Shape CastingDocument3 pagesNear Net Shape CastingNURNo ratings yet
- F. D. M. Haldane PDFDocument4 pagesF. D. M. Haldane PDFHamid iqbalNo ratings yet
- Autoclave Reactor PatentDocument9 pagesAutoclave Reactor PatentPrabandari 632No ratings yet
- PDFDocument28 pagesPDFMiraNo ratings yet
- Android Development For C# and Visual Studio 2012Document2 pagesAndroid Development For C# and Visual Studio 2012johnvhorcajoNo ratings yet
- How To Draw and Analyse Free Body Diagrams (FBDS) - Ilearn Engineering®Document20 pagesHow To Draw and Analyse Free Body Diagrams (FBDS) - Ilearn Engineering®wajju007No ratings yet
- Mechanics of Solid (Mac11) Assignment: When Tensile Stress Is Applied Axially On A Circular Rod ItsDocument3 pagesMechanics of Solid (Mac11) Assignment: When Tensile Stress Is Applied Axially On A Circular Rod ItsAyush SharmaNo ratings yet
- We Choose To Go To The Moon Speech by John F. Kennedy September 12th 1962Document4 pagesWe Choose To Go To The Moon Speech by John F. Kennedy September 12th 1962ViAnne ReyesNo ratings yet
- Instruction Manual S 400Document29 pagesInstruction Manual S 400Sơn Lê CaoNo ratings yet
- GS Cold Mix Process For Blended Spreads GBDocument8 pagesGS Cold Mix Process For Blended Spreads GBNicolas BenavidezNo ratings yet
- Internship Report Kandhkot Gas Field: Submitted byDocument10 pagesInternship Report Kandhkot Gas Field: Submitted byMuzzamilNo ratings yet
- Lindab Round Duct and Fitting PDFDocument108 pagesLindab Round Duct and Fitting PDFLinh TruongNo ratings yet
- Technical Paper - CIGREDocument12 pagesTechnical Paper - CIGREGobinath BalasubramanianNo ratings yet
- O2 and A/F Sensor Diagnosis: Section 7Document44 pagesO2 and A/F Sensor Diagnosis: Section 7sungjoo75No ratings yet
- Lou's Approved Vendor List: Service Company Name PhoneDocument2 pagesLou's Approved Vendor List: Service Company Name Phonecutty54No ratings yet
- Iso 1461-2022Document22 pagesIso 1461-2022Huipeng li100% (1)
- SVANPC Plusplus 2 0Document221 pagesSVANPC Plusplus 2 0Basilio DanziNo ratings yet
- Vendor Component Libraries - RF Passive SMT LibraryDocument114 pagesVendor Component Libraries - RF Passive SMT LibraryKyusang ParkNo ratings yet
- Min To Wheel Calculation UpdateDocument16 pagesMin To Wheel Calculation UpdateterryltharpNo ratings yet
- Engine Overall Engine Specification Engine Proper Valve Mechanism Cooling System Fuel System Ignition System Engine Mounting Engine Control SystemDocument40 pagesEngine Overall Engine Specification Engine Proper Valve Mechanism Cooling System Fuel System Ignition System Engine Mounting Engine Control SystemPurnama Art100% (2)
- Seaton Mulberry Homes Floor PlansDocument34 pagesSeaton Mulberry Homes Floor PlansPicchigaNo ratings yet
- 1404 E EUC 3 Manual and Automatic Compensation of Reactive PowerDocument20 pages1404 E EUC 3 Manual and Automatic Compensation of Reactive PowerРаденко МарјановићNo ratings yet
- Saic D 2025Document10 pagesSaic D 2025jerinNo ratings yet
- Datasheet of DS-2CD1021-I (D) V5.5.4 20180313Document4 pagesDatasheet of DS-2CD1021-I (D) V5.5.4 20180313diego lopezNo ratings yet
- Aerodynamic 1Document6 pagesAerodynamic 1BOBNo ratings yet
- Br-PrestressedBeam V31jan19 1Document9 pagesBr-PrestressedBeam V31jan19 1Rikesh SapkotaNo ratings yet
- Encapsulation, ADT, Inheritance and PolymorphismpptDocument20 pagesEncapsulation, ADT, Inheritance and Polymorphismpptdivya_167No ratings yet
- Research Paper #1Document38 pagesResearch Paper #1Meynard Orbino33% (3)