Professional Documents
Culture Documents
Heart Diesese
Heart Diesese
Uploaded by
Suhani ShindeOriginal Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Heart Diesese
Heart Diesese
Uploaded by
Suhani ShindeCopyright:
Available Formats
4/15/24, 8:40 AM Heart diesese
In [1]: # import pandas library
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import accuracy_score,confusion_matrix
from sklearn.linear_model import LogisticRegression
import seaborn as sns
import matplotlib.pyplot as plt
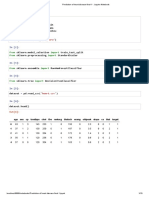
In [2]: # Reading csv file
df = pd.read_csv("Heart.csv")
df.head()
Out[2]: age sex cp trtbps chol fbs restecg thalachh exng oldpeak slp caa thall output
0 63 1 3 145 233 1 0 150 0 2.3 0 0 1 1
1 37 1 2 130 250 0 1 187 0 3.5 0 0 2 1
2 41 0 1 130 204 0 0 172 0 1.4 2 0 2 1
3 56 1 1 120 236 0 1 178 0 0.8 2 0 2 1
4 57 0 0 120 354 0 1 163 1 0.6 2 0 2 1
Data Cleaning
In [3]: df = df.drop_duplicates()
In [4]: # Count ,min,max ,etc of each column
df.describe()
Out[4]: age sex cp trtbps chol fbs restecg thalac
count 302.00000 302.000000 302.000000 302.000000 302.000000 302.000000 302.000000 302.0000
mean 54.42053 0.682119 0.963576 131.602649 246.500000 0.149007 0.526490 149.5695
std 9.04797 0.466426 1.032044 17.563394 51.753489 0.356686 0.526027 22.9035
min 29.00000 0.000000 0.000000 94.000000 126.000000 0.000000 0.000000 71.0000
25% 48.00000 0.000000 0.000000 120.000000 211.000000 0.000000 0.000000 133.2500
50% 55.50000 1.000000 1.000000 130.000000 240.500000 0.000000 1.000000 152.5000
75% 61.00000 1.000000 2.000000 140.000000 274.750000 0.000000 1.000000 166.0000
max 77.00000 1.000000 3.000000 200.000000 564.000000 1.000000 2.000000 202.0000
In [5]: # Information about each column data
df.info()
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 1/9
4/15/24, 8:40 AM Heart diesese
<class 'pandas.core.frame.DataFrame'>
Index: 302 entries, 0 to 302
Data columns (total 14 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 age 302 non-null int64
1 sex 302 non-null int64
2 cp 302 non-null int64
3 trtbps 302 non-null int64
4 chol 302 non-null int64
5 fbs 302 non-null int64
6 restecg 302 non-null int64
7 thalachh 302 non-null int64
8 exng 302 non-null int64
9 oldpeak 302 non-null float64
10 slp 302 non-null int64
11 caa 302 non-null int64
12 thall 302 non-null int64
13 output 302 non-null int64
dtypes: float64(1), int64(13)
memory usage: 35.4 KB
In [6]: #Finding null values in each column
df.isna().sum()
age 0
Out[6]:
sex 0
cp 0
trtbps 0
chol 0
fbs 0
restecg 0
thalachh 0
exng 0
oldpeak 0
slp 0
caa 0
thall 0
output 0
dtype: int64
Data Integration
In [7]: df.head()
Out[7]: age sex cp trtbps chol fbs restecg thalachh exng oldpeak slp caa thall output
0 63 1 3 145 233 1 0 150 0 2.3 0 0 1 1
1 37 1 2 130 250 0 1 187 0 3.5 0 0 2 1
2 41 0 1 130 204 0 0 172 0 1.4 2 0 2 1
3 56 1 1 120 236 0 1 178 0 0.8 2 0 2 1
4 57 0 0 120 354 0 1 163 1 0.6 2 0 2 1
In [8]: df.fbs.unique()
array([1, 0], dtype=int64)
Out[8]:
In [9]: subSet1 = df[['age','cp','chol','thalachh']]
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 2/9
4/15/24, 8:40 AM Heart diesese
In [10]: subSet2 = df[['exng','slp','output']]
In [11]: merged_df = subSet1.merge(right=subSet2,how='cross')
merged_df.head()
Out[11]: age cp chol thalachh exng slp output
0 63 3 233 150 0 0 1
1 63 3 233 150 0 0 1
2 63 3 233 150 0 2 1
3 63 3 233 150 0 2 1
4 63 3 233 150 1 2 1
Error Correcting
In [12]: df.columns
Index(['age', 'sex', 'cp', 'trtbps', 'chol', 'fbs', 'restecg', 'thalachh',
Out[12]:
'exng', 'oldpeak', 'slp', 'caa', 'thall', 'output'],
dtype='object')
In [13]: def remove_outliers(column):
Q1 = column.quantile(0.25)
Q3 = column.quantile(0.75)
IQR = Q3 - Q1
threshold = 1.5 * IQR
outlier_mask = (column < Q1 - threshold) | (column > Q3 + threshold)
return column[~outlier_mask]
In [14]: # Remove outliers for each column using a loop
col_name = ['cp','thalachh','exng','oldpeak','slp','caa']
for col in col_name:
df[col] = remove_outliers(df[col])
In [15]: plt.figure(figsize=(10, 6)) # Adjust the figure size if needed
for col in col_name:
sns.boxplot(data=df[col])
plt.title(col)
plt.show()
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 3/9
4/15/24, 8:40 AM Heart diesese
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 4/9
4/15/24, 8:40 AM Heart diesese
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 5/9
4/15/24, 8:40 AM Heart diesese
In [16]: df = df.dropna()
In [17]: df.isna().sum()
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 6/9
4/15/24, 8:40 AM Heart diesese
age 0
Out[17]:
sex 0
cp 0
trtbps 0
chol 0
fbs 0
restecg 0
thalachh 0
exng 0
oldpeak 0
slp 0
caa 0
thall 0
output 0
dtype: int64
In [18]: df = df.drop('fbs',axis=1)
In [19]: # Compute correlations between features and target
correlations = df.corr()['output'].drop('output')
# Print correlations
print("Correlation with the Target:")
print(correlations)
print()
# Plot correlation heatmap
plt.figure(figsize=(8, 6))
sns.heatmap(df.corr(), annot=True, cmap='coolwarm')
plt.title('Correlation Heatmap')
plt.show()
Correlation with the Target:
age -0.193798
sex -0.303271
cp 0.410807
trtbps -0.135238
chol -0.052796
restecg 0.122071
thalachh 0.384609
exng -0.444401
oldpeak -0.437895
slp 0.329432
caa -0.460816
thall -0.366390
Name: output, dtype: float64
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 7/9
4/15/24, 8:40 AM Heart diesese
In [20]: # df.isna().sum()
Data Split
In [21]: # splitting data using train test split
x = df[['cp','thalachh','exng','oldpeak','slp','caa']]
y = df.output
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.2,random_state=0)
x_train.shape,x_test.shape,y_train.shape,y_test.shape
((220, 6), (55, 6), (220,), (55,))
Out[21]:
Data transformation
In [22]: from sklearn.preprocessing import StandardScaler
In [23]: scaler = StandardScaler()
In [24]: x_train_scaled = scaler.fit_transform(x_train)
x_test_scaled = scaler.transform(x_test)
In [ ]:
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 8/9
4/15/24, 8:40 AM Heart diesese
Data model building
In [25]: y_train= np.array(y_train).reshape(-1, 1)
y_test= np.array(y_test).reshape(-1, 1)
In [26]: y_train.shape
(220, 1)
Out[26]:
In [27]: model = LogisticRegression()
model.fit(x_train_scaled, y_train)
# Make predictions on the test set
y_pred = model.predict(x_test_scaled)
# Evaluate the model's accuracy
accuracy = accuracy_score(y_test, y_pred)
print("Accuracy:", accuracy)
Accuracy: 0.8363636363636363
C:\Users\Admin\anaconda3\Lib\site-packages\sklearn\utils\validation.py:1184: DataC
onversionWarning: A column-vector y was passed when a 1d array was expected. Pleas
e change the shape of y to (n_samples, ), for example using ravel().
y = column_or_1d(y, warn=True)
In [28]: #Classification model using Decision Tree
from sklearn.tree import DecisionTreeClassifier
tc=DecisionTreeClassifier(criterion='entropy')
tc.fit(x_train_scaled,y_train)
y_pred=tc.predict(x_test_scaled)
print("Training Accuracy Score :",accuracy_score(y_pred,y_test))
print("Training Confusion Matrix :",confusion_matrix(y_pred,y_test))
Training Accuracy Score : 0.8363636363636363
Training Confusion Matrix : [[21 3]
[ 6 25]]
In [ ]:
localhost:8888/nbconvert/html/Desktop/DSBDA GROP B/Assignment 2/Heart diesese.ipynb?download=false 9/9
You might also like
- 1.1 Read The Data and Do Exploratory Data Analysis. Describe The Data BrieflyDocument50 pages1.1 Read The Data and Do Exploratory Data Analysis. Describe The Data BrieflyP Venkata Krishna Rao100% (19)
- Bizhub Pro 951 Parts ManualDocument262 pagesBizhub Pro 951 Parts Manualdmcampos63% (8)
- Management Advisory Services by Roque Solution Manual PDFDocument3 pagesManagement Advisory Services by Roque Solution Manual PDFLegara Robelyn25% (28)
- Java: The Complete Reference, Eleventh Edition - Herbert SchildtDocument5 pagesJava: The Complete Reference, Eleventh Edition - Herbert Schildtzykohyba0% (3)
- Virtual Lab Stopping Distance of A CarDocument4 pagesVirtual Lab Stopping Distance of A Carrdixit20% (1)
- 52 MX ErdDocument123 pages52 MX Erdacha847No ratings yet
- Sentimental Analysis Project DocumentationDocument67 pagesSentimental Analysis Project DocumentationSukanya Vemulapalli75% (4)
- ML Project - End-To-End-Heart-Disease-ClassificationDocument30 pagesML Project - End-To-End-Heart-Disease-ClassificationAnurag KumarNo ratings yet
- QUIZ Week 2 CART Practice PDFDocument10 pagesQUIZ Week 2 CART Practice PDFNagarajan ThandayuthamNo ratings yet
- Heart Dataset AnalysisDocument24 pagesHeart Dataset Analysiseseoghene emebraduNo ratings yet
- Heart: Our "Goal" Predict The Presence of Heart Disease in The PatientDocument73 pagesHeart: Our "Goal" Predict The Presence of Heart Disease in The Patientaditya b100% (1)
- B 4 HeartDocument9 pagesB 4 HeartSuhani ShindeNo ratings yet
- Prediction of Heart Disease Final-1-20 April - Jupyter NotebookDocument10 pagesPrediction of Heart Disease Final-1-20 April - Jupyter NotebookRushabh JainNo ratings yet
- Logistic RegressionDocument12 pagesLogistic RegressionKagade AjinkyaNo ratings yet
- Heart Failure PredictionDocument41 pagesHeart Failure PredictionAKKALA VIJAYGOUD100% (1)
- Ide To 6 Classification AlgorithmsDocument34 pagesIde To 6 Classification AlgorithmsAssy GlendaNo ratings yet
- Python SolutionDocument30 pagesPython SolutionMile MileNo ratings yet
- Heart Attacks AnalysisDocument10 pagesHeart Attacks AnalysisNarasimhaNo ratings yet
- Seaborn - Plots - Jupyter NotebookDocument36 pagesSeaborn - Plots - Jupyter NotebookLahari BanalaNo ratings yet
- B - 59 - SMA - Exp 4Document9 pagesB - 59 - SMA - Exp 4Ritz FernandesNo ratings yet
- Dsbda 4Document4 pagesDsbda 4Arbaz ShaikhNo ratings yet
- Correlation: Import As Import As Import As Import As From Import From Import Import Matplotlib ImportDocument1 pageCorrelation: Import As Import As Import As Import As From Import From Import Import Matplotlib ImportHarsh BangiaNo ratings yet
- Users of A Music Streaming Service Will Churn or Stay: @staticmethodDocument1 pageUsers of A Music Streaming Service Will Churn or Stay: @staticmethodGowtham SuguNo ratings yet
- Import As Import As: "Iris - CSV"Document4 pagesImport As Import As: "Iris - CSV"avinashNo ratings yet
- Id Sepallengthcm Sepalwidthcm Petallengthcm Petalwidthcm Species 0 1 2 3 4Document4 pagesId Sepallengthcm Sepalwidthcm Petallengthcm Petalwidthcm Species 0 1 2 3 4Sebastian CallesNo ratings yet
- Logistic Regression 205Document8 pagesLogistic Regression 205Ranadeep DeyNo ratings yet
- Dsbda 10Document3 pagesDsbda 10monaliauti2No ratings yet
- Heart Disease Prediction - Jupyter NotebookDocument9 pagesHeart Disease Prediction - Jupyter Notebookkavya100% (1)
- 20MIS1025 PerceptronDocument5 pages20MIS1025 PerceptronSandip DasNo ratings yet
- Week - 6 - SWI - MLP - LogisticRegression - Ipynb - ColaboratoryDocument15 pagesWeek - 6 - SWI - MLP - LogisticRegression - Ipynb - ColaboratoryMeer HassanNo ratings yet
- Linear - Regression - Ipynb - ColaboratoryDocument4 pagesLinear - Regression - Ipynb - Colaboratoryavnimote121No ratings yet
- Satya772244@gmail - Com House Price PredictionDocument5 pagesSatya772244@gmail - Com House Price PredictionSatyendra VermaNo ratings yet
- Assign9.Ipynb - ColabDocument4 pagesAssign9.Ipynb - Colabatharvapudale0608No ratings yet
- Observation: As We Can See We Have Threwe Types of Datatypes I.E. (Int, Float, Object) That Means We Have Both Categorical and Numerical DataDocument2 pagesObservation: As We Can See We Have Threwe Types of Datatypes I.E. (Int, Float, Object) That Means We Have Both Categorical and Numerical DataPRATIK RATHODNo ratings yet
- Exp 4 (Movie Review)Document7 pagesExp 4 (Movie Review)pameluftNo ratings yet
- Lab11 Inlab and PostlabDocument5 pagesLab11 Inlab and PostlabHelloNo ratings yet
- Introduction To Scikit LearnDocument108 pagesIntroduction To Scikit LearnRaiyan Zannat100% (1)
- Assignment3 VidulGargDocument14 pagesAssignment3 VidulGargvidulgarg1524No ratings yet
- 7-8 Feature Engineering 101-NormalizationDocument8 pages7-8 Feature Engineering 101-NormalizationKhan RafiNo ratings yet
- Parcial2-Javier CardenasDocument7 pagesParcial2-Javier CardenasJavier CardenasNo ratings yet
- Income Qualification Project3Document40 pagesIncome Qualification Project3NikhilNo ratings yet
- Loadalgarve MLPDocument7 pagesLoadalgarve MLPRony RockNo ratings yet
- 04 BigqueryDocument6 pages04 BigqueryjenNo ratings yet
- Hasil RendiDocument11 pagesHasil RendiFitria KintaNo ratings yet
- Instructions:: Mltest2question - Jupyter NotebookDocument6 pagesInstructions:: Mltest2question - Jupyter NotebookAniruddha TrivediNo ratings yet
- Import As From Import From Import From Import From Import From Import From Import From Import From Import From Import From Import Import AsDocument8 pagesImport As From Import From Import From Import From Import From Import From Import From Import From Import From Import From Import Import Asvince.lachicaNo ratings yet
- RobiSetiawan - Tugas 4 .IpynbDocument13 pagesRobiSetiawan - Tugas 4 .IpynbROBI SETIAWANNo ratings yet
- Haberman Datasets Analysis - Ipynb - ColaboratoryDocument13 pagesHaberman Datasets Analysis - Ipynb - ColaboratoryShyamal HazarikaNo ratings yet
- Sunbase Data AssignmentDocument11 pagesSunbase Data Assignmentfanrock281No ratings yet
- Practical-5 - Jupyter NotebookDocument8 pagesPractical-5 - Jupyter NotebookHarsha Gohil100% (1)
- Logistic - Ipynb - ColaboratoryDocument6 pagesLogistic - Ipynb - ColaboratoryAkansha UniyalNo ratings yet
- DL (Pra 01)Document9 pagesDL (Pra 01)Dhanashri SalunkheNo ratings yet
- AidsDocument88 pagesAidsCarlos Huerta SalasNo ratings yet
- Covid-19 Prediction - Jupyter NotebookDocument6 pagesCovid-19 Prediction - Jupyter NotebookTEEGALAMEHERRITHWIK 122010320035No ratings yet
- Practica 11Document7 pagesPractica 112marlenehh2003No ratings yet
- Output1Document8 pagesOutput1Laptop-Dimas-249No ratings yet
- Decision Tree Algorithm 1677030641Document9 pagesDecision Tree Algorithm 1677030641juanNo ratings yet
- Lab 3. Linear Regression 230223Document7 pagesLab 3. Linear Regression 230223ruso100% (1)
- Heart Disease Classification ML Assignment - Jupyter NotebookDocument7 pagesHeart Disease Classification ML Assignment - Jupyter Notebookfurole sammyceNo ratings yet
- Decision TreeDocument3 pagesDecision Tree20CSE102 SUDHARSAN KNo ratings yet
- Ash 4Document8 pagesAsh 4saurabhbborate0621No ratings yet
- Decision Tree Regressor and Ensemble Techniques - RegressorsDocument18 pagesDecision Tree Regressor and Ensemble Techniques - RegressorsS ANo ratings yet
- Assign10.Ipynb - ColabDocument8 pagesAssign10.Ipynb - Colabatharvapudale0608No ratings yet
- Statistical Data Analysis - Ipynb - ColaboratoryDocument6 pagesStatistical Data Analysis - Ipynb - ColaboratoryVarad KulkarniNo ratings yet
- Logistic Pima Indians - Ipynb - ColaboratoryDocument4 pagesLogistic Pima Indians - Ipynb - ColaboratorySHEKHAR SWAMINo ratings yet
- Cable Tray Management Catalog - BLINE COOPERDocument454 pagesCable Tray Management Catalog - BLINE COOPERInVisBLNo ratings yet
- Topica-500-User Timer 03Document1 pageTopica-500-User Timer 03eeindustrialNo ratings yet
- Alarm CommlistDocument168 pagesAlarm CommlistSyahrul WahyudiNo ratings yet
- Algebra 1 Scope and Sequence 2018-2019Document21 pagesAlgebra 1 Scope and Sequence 2018-2019api-283114543100% (1)
- Veenet Technological Service - E BrochureDocument3 pagesVeenet Technological Service - E BrochureRam PrakashNo ratings yet
- 4.2 The Principles of Modern Software Management 63Document4 pages4.2 The Principles of Modern Software Management 63Deepak KumarNo ratings yet
- Comparing RGB-contone - CMYK Contone and Halftone Printer Drivers PDFDocument4 pagesComparing RGB-contone - CMYK Contone and Halftone Printer Drivers PDFFelix Alfredo Vilchez TupayachiNo ratings yet
- Embedded Systems and Information AppliancesDocument11 pagesEmbedded Systems and Information Appliancesshaikshaa007100% (4)
- Index: LTE Signaling, Troubleshooting and Optimization, First Edition. Ralf Kreher and Karsten GaengerDocument6 pagesIndex: LTE Signaling, Troubleshooting and Optimization, First Edition. Ralf Kreher and Karsten Gaengersuminda456No ratings yet
- Galaxy G1 User ManualDocument70 pagesGalaxy G1 User ManualRandy Mucha V.No ratings yet
- Technical Interview Feedback Form: Online Assessment ScoreDocument36 pagesTechnical Interview Feedback Form: Online Assessment ScoreTokala Rajesh0% (1)
- Davian Litcmore CAPE Comp Sci Unit2 Ia 2011Document80 pagesDavian Litcmore CAPE Comp Sci Unit2 Ia 2011Davian Kal'El LitchmoreNo ratings yet
- Lecture 19 Maple 2017Document101 pagesLecture 19 Maple 2017Stephen AlaoNo ratings yet
- General Guidelines For Successful Materials SelectionDocument5 pagesGeneral Guidelines For Successful Materials SelectionBALARAM S PATTARNo ratings yet
- A Novel Approach To Estimate The Gap Between The Middle and Short-Term PlansDocument8 pagesA Novel Approach To Estimate The Gap Between The Middle and Short-Term PlansConiCortesLizamaNo ratings yet
- Debian GNU/Linux Installation GuideDocument130 pagesDebian GNU/Linux Installation Guideopenid_jo0Lc0b4No ratings yet
- HP NonStop TACL Reference ManualDocument950 pagesHP NonStop TACL Reference ManualAjay100% (1)
- Cyber-Ark Password VaultDocument14 pagesCyber-Ark Password VaultoliveNo ratings yet
- Module 1: Introducing Application Lifecycle ManagementDocument28 pagesModule 1: Introducing Application Lifecycle Managementeduardo_quintanill_3No ratings yet
- Exercise 1 & 2Document14 pagesExercise 1 & 2jasmineNo ratings yet
- CS153: Compilers Lecture 19: Optimization: Stephen ChongDocument38 pagesCS153: Compilers Lecture 19: Optimization: Stephen Chongkavya sri gNo ratings yet
- RubricDocument4 pagesRubricMuhammad Irfan MalikNo ratings yet
- Mapping and Characterization of Rubber Plantations Using Resourcesat DataDocument1 pageMapping and Characterization of Rubber Plantations Using Resourcesat DatapradeepNo ratings yet
- Write A Program To Store The Elements in 1-D Array and Perform The Operations Like Searching, Sorting and Reversing The Elements. (Menu Driven)Document11 pagesWrite A Program To Store The Elements in 1-D Array and Perform The Operations Like Searching, Sorting and Reversing The Elements. (Menu Driven)BalavantNo ratings yet