Professional Documents
Culture Documents
(W-6302) Neural Mode Jump Monte Carlo
(W-6302) Neural Mode Jump Monte Carlo
Uploaded by
Nazmul SaqibCopyright:
Available Formats
You might also like
- College Physics 4th Edition Giambattista Test Bank 1Document31 pagesCollege Physics 4th Edition Giambattista Test Bank 1tracy100% (43)
- An Extension of Newton-Type Algorithms For Nonlinear Process ControlDocument6 pagesAn Extension of Newton-Type Algorithms For Nonlinear Process ControlGustavo Gabriel Jimenez100% (1)
- 220-12 Volt DC Power SupplyDocument16 pages220-12 Volt DC Power Supplychachunasayan89% (18)
- Temperature Dependence of Energy Transport in The Z Chiral Clock ModelDocument12 pagesTemperature Dependence of Energy Transport in The Z Chiral Clock ModelDaniele MarconiNo ratings yet
- Project ProposalDocument10 pagesProject ProposalPrathamNo ratings yet
- Andreas M. Lauchli - Numerical Simulations of Frustrated SystemsDocument34 pagesAndreas M. Lauchli - Numerical Simulations of Frustrated SystemsImaxSWNo ratings yet
- ChuangDocument8 pagesChuangKamal ChoudharyNo ratings yet
- Machine Learning Phases of Strongly Correlated FermionsDocument9 pagesMachine Learning Phases of Strongly Correlated FermionsalfonsoNo ratings yet
- King Et Al-2021-Nature CommunicationsDocument6 pagesKing Et Al-2021-Nature CommunicationsSteve KingNo ratings yet
- A Reduced Variational Approach For Searching Cycles in High-Dimensional SystemsDocument25 pagesA Reduced Variational Approach For Searching Cycles in High-Dimensional Systemsim.marzaNo ratings yet
- Meshless Methods1Document4 pagesMeshless Methods1herevenkatNo ratings yet
- Recognition of Temporal Sequences of Patterns Using State-Dependent SynapsesDocument18 pagesRecognition of Temporal Sequences of Patterns Using State-Dependent Synapsesyixinah516No ratings yet
- Entire Spectrum of Fully Many-Body Localized Systems Using Tensor NetworksDocument12 pagesEntire Spectrum of Fully Many-Body Localized Systems Using Tensor NetworksAnonymous lqvgUsVckNo ratings yet
- 1 s2.0 S0370269323004227 MainDocument7 pages1 s2.0 S0370269323004227 Mainsakoufah adrianNo ratings yet
- Reese JM Pure Coupled Continuum Hydrodynamics and Molecular Dynamics Method For Multiscale Simulation Aug 2011Document9 pagesReese JM Pure Coupled Continuum Hydrodynamics and Molecular Dynamics Method For Multiscale Simulation Aug 2011Avijit KarmakarNo ratings yet
- Clementi Diffussion MAps JCPDocument11 pagesClementi Diffussion MAps JCPDavid BeveridgeNo ratings yet
- Hard-Disk Equation of State: First-Order Liquid-Hexatic Transition in Two Dimensions With Three Simulation MethodsDocument8 pagesHard-Disk Equation of State: First-Order Liquid-Hexatic Transition in Two Dimensions With Three Simulation MethodsOzanBurakEriçokNo ratings yet
- Physical Quantum AlgorithmsDocument7 pagesPhysical Quantum AlgorithmsSankha DeepNo ratings yet
- Non-Markovian Quantum Dynamics in Strongly Coupled Multimode Cavities Conditioned On Continuous MeasurementDocument16 pagesNon-Markovian Quantum Dynamics in Strongly Coupled Multimode Cavities Conditioned On Continuous MeasurementMartta TervakoskiNo ratings yet
- 1104 5491v2 PDFDocument19 pages1104 5491v2 PDFJacky LaoNo ratings yet
- Quantum Monte CarloDocument6 pagesQuantum Monte Carlomar.sedanoortNo ratings yet
- Laurence E. Fried and Gregory S. Ezra - PERTURB: A Special-Purpose Algebraic Manipulation Program For Classical Perturbation TheoryDocument15 pagesLaurence E. Fried and Gregory S. Ezra - PERTURB: A Special-Purpose Algebraic Manipulation Program For Classical Perturbation TheoryOmasazzNo ratings yet
- Nino Garis - Numerical Solutions of The One Speed Neutron Transport Equation in Two Medium Slabs and Spheres (1991)Document52 pagesNino Garis - Numerical Solutions of The One Speed Neutron Transport Equation in Two Medium Slabs and Spheres (1991)Strawberry FieldsNo ratings yet
- RadevDocument3 pagesRadevIrenataNo ratings yet
- Umbrella Sampling - A ReviewDocument11 pagesUmbrella Sampling - A ReviewdanieloshkaNo ratings yet
- Decoherence ProblemDocument6 pagesDecoherence ProblemmrpcuNo ratings yet
- Spontaneous Symmetry Breaking in Replica Field Theory: 10.1103/physrevd.96.065012Document14 pagesSpontaneous Symmetry Breaking in Replica Field Theory: 10.1103/physrevd.96.065012Angie DamarisNo ratings yet
- Andrew K. Henrick, Tariq D. Aslam and Joseph M. Powers - Simulations of Pulsating One-Dimensional Detonations With True Fifth Order AccuracyDocument19 pagesAndrew K. Henrick, Tariq D. Aslam and Joseph M. Powers - Simulations of Pulsating One-Dimensional Detonations With True Fifth Order AccuracyNikeShoxxxNo ratings yet
- 1609 01552 PDFDocument12 pages1609 01552 PDFGNo ratings yet
- 0907 2948 PDFDocument17 pages0907 2948 PDFCarlos LopezNo ratings yet
- Certifying Multilevel Coherence in The Motional State of A Trapped IonDocument14 pagesCertifying Multilevel Coherence in The Motional State of A Trapped IonNikhilKotibhaskarNo ratings yet
- Data-Driven Computation of Molecular Reaction Coordinates: Andreas Bittracher, Ralf Banisch, and Christof SCH UtteDocument24 pagesData-Driven Computation of Molecular Reaction Coordinates: Andreas Bittracher, Ralf Banisch, and Christof SCH Uttehuevonomar05No ratings yet
- Boost-Invariant (2+1) - Dimensional Anisotropic Hydrodynamics: PACS Numbers: 12.38.Mh, 24.10.Nz, 25.75.Ld, 25.75.-qDocument44 pagesBoost-Invariant (2+1) - Dimensional Anisotropic Hydrodynamics: PACS Numbers: 12.38.Mh, 24.10.Nz, 25.75.Ld, 25.75.-qsharky84No ratings yet
- Decoherence Limits The Cost To Simulate An Anharmonic OscillatorDocument21 pagesDecoherence Limits The Cost To Simulate An Anharmonic OscillatorSRINIDHI PAWARNo ratings yet
- Numerical Atomic Orbitals For Linear-Scaling Calculations: 共Received 6 April 2001; Published 28 November 2001兲Document9 pagesNumerical Atomic Orbitals For Linear-Scaling Calculations: 共Received 6 April 2001; Published 28 November 2001兲AytaçÇelikNo ratings yet
- Distributed Quantum Phase Estimation With Entangled PhotonsDocument8 pagesDistributed Quantum Phase Estimation With Entangled Photons박정우No ratings yet
- Phase Equilibria in The Polydisperse Zwanzig Model of Hard RodsDocument14 pagesPhase Equilibria in The Polydisperse Zwanzig Model of Hard RodsGilberto RuizNo ratings yet
- Sequence Space JacobianDocument84 pagesSequence Space JacobianStan ChrisinskyNo ratings yet
- QMMM MethodDocument8 pagesQMMM MethodSubarnaNo ratings yet
- Load Balancing Strategies For The DSMC Simulation of Hypersonic Flows Using HPCDocument12 pagesLoad Balancing Strategies For The DSMC Simulation of Hypersonic Flows Using HPCnanilaurntiuNo ratings yet
- Modelos DFTDocument17 pagesModelos DFTAlberto Núñez CardezoNo ratings yet
- Non Linear CH PaperDocument9 pagesNon Linear CH PaperseenuNo ratings yet
- Statement of Research Interests-Example 3Document7 pagesStatement of Research Interests-Example 3NedelcuGeorgeNo ratings yet
- Collective Variable-Based Enhanced Sampling and Machine Learning PDFDocument17 pagesCollective Variable-Based Enhanced Sampling and Machine Learning PDFwenhuishenNo ratings yet
- Nonlinear Heat Transport in Mesoscopic Conductors: Rectification, Peltier Effect and Wiedemann-Franz LawDocument8 pagesNonlinear Heat Transport in Mesoscopic Conductors: Rectification, Peltier Effect and Wiedemann-Franz LawJeanette AmayaNo ratings yet
- An Adaptive Variational Algorithm For Exact Molecular Simulations On A Quantum ComputerDocument9 pagesAn Adaptive Variational Algorithm For Exact Molecular Simulations On A Quantum ComputerRaiyan RahmanNo ratings yet
- 07 - chapter 2 مهم جدا جدا لحساب activation energyDocument30 pages07 - chapter 2 مهم جدا جدا لحساب activation energymahmoudNo ratings yet
- Micadei - Experimental Rectification of Entropy Production by A Maxwell's Demon in A Quantum SystemDocument10 pagesMicadei - Experimental Rectification of Entropy Production by A Maxwell's Demon in A Quantum SystemFábio LucasNo ratings yet
- Numerical Algorithms For The Hypercube Concu - 1988 - Mathematical and ComputerDocument3 pagesNumerical Algorithms For The Hypercube Concu - 1988 - Mathematical and ComputerdarshanNo ratings yet
- The Emergent Copenhagen Interpretation of Quantum Mechanics - Timothy J. HollowoodDocument21 pagesThe Emergent Copenhagen Interpretation of Quantum Mechanics - Timothy J. HollowoodCambiador de MundoNo ratings yet
- Coherent Feedback Cooling of A Nanomechanical Membrane With Atomic SpinsDocument13 pagesCoherent Feedback Cooling of A Nanomechanical Membrane With Atomic Spinswork outNo ratings yet
- GPU-based Reaction Ensemble Monte Carlo Method For Non-Ideal Plasma ThermodynamicsDocument28 pagesGPU-based Reaction Ensemble Monte Carlo Method For Non-Ideal Plasma ThermodynamicsAndreina Dell'olioNo ratings yet
- Mesoscopic Quantum Coherences in Cavity QED: Preparation and Decoherence Monitoring SchemesDocument15 pagesMesoscopic Quantum Coherences in Cavity QED: Preparation and Decoherence Monitoring SchemesVictor Viera CastilloNo ratings yet
- 123Document4 pages123vato otavNo ratings yet
- On The Mathematical Foundations of Diffusion Monte Carlo: Michel Caffarel, Pierre Del Moral and Luc de MontellaDocument38 pagesOn The Mathematical Foundations of Diffusion Monte Carlo: Michel Caffarel, Pierre Del Moral and Luc de MontellaPinakiNo ratings yet
- E (3) - Equivariant Graph Neural Networks For Data-Ef Ficient and Accurate Interatomic PotentialsDocument11 pagesE (3) - Equivariant Graph Neural Networks For Data-Ef Ficient and Accurate Interatomic PotentialsichbinkeinrobotjaNo ratings yet
- Paper On Umbrella SamplingDocument11 pagesPaper On Umbrella SamplingSayed Abu SufyanNo ratings yet
- Mathematics 09 01442Document12 pagesMathematics 09 01442Golam ZakariaNo ratings yet
- (1959) Studies in Molecular Dynamics. I. General MethodDocument8 pages(1959) Studies in Molecular Dynamics. I. General MethodJakler NicheleNo ratings yet
- Polarization Coding of Quantum Information (Optics Communications, Vol. 123, Issue 1-3) (1996)Document10 pagesPolarization Coding of Quantum Information (Optics Communications, Vol. 123, Issue 1-3) (1996)sepot24093No ratings yet
- Rodrguez Her N Ndez 2014Document6 pagesRodrguez Her N Ndez 2014Tengku Arief Buana PerkasaNo ratings yet
- Journal of Quantitative Spectroscopy & Radiative Transfer: Laurent Soucasse, Philippe Riviere, Anouar SoufianiDocument9 pagesJournal of Quantitative Spectroscopy & Radiative Transfer: Laurent Soucasse, Philippe Riviere, Anouar Soufianimbb appNo ratings yet
- (W-7443) Differentiable Molecular Simulations For Control and LearningDocument16 pages(W-7443) Differentiable Molecular Simulations For Control and LearningNazmul SaqibNo ratings yet
- (W-6135) (2023) Accelerated Sampling of Rare Events Using A Neural Network Bias PotentialDocument12 pages(W-6135) (2023) Accelerated Sampling of Rare Events Using A Neural Network Bias PotentialNazmul SaqibNo ratings yet
- (W-6725) TensorMol - NN Augmented With Long Ramge PhysicsDocument9 pages(W-6725) TensorMol - NN Augmented With Long Ramge PhysicsNazmul SaqibNo ratings yet
- (W-6597) Deep-Neural-Network Approach To Solving The Ab Initio Nuclear Structure ProblemDocument9 pages(W-6597) Deep-Neural-Network Approach To Solving The Ab Initio Nuclear Structure ProblemNazmul SaqibNo ratings yet
- How To Do Machine Learning With Small DataDocument22 pagesHow To Do Machine Learning With Small DataNazmul SaqibNo ratings yet
- Series EA Electric Actuators Low Torque, Medium Torque, and Spring Return Manual 1321-In-003!0!13Document16 pagesSeries EA Electric Actuators Low Torque, Medium Torque, and Spring Return Manual 1321-In-003!0!13Isaac MonterreyNo ratings yet
- Arches Vaults and Dome FinalDocument39 pagesArches Vaults and Dome FinalPratima Mahesh0% (1)
- 096 09247 03 - TDSDocument2 pages096 09247 03 - TDSMuhamad Samsul BahriNo ratings yet
- WCM - Yamashina - 05 - Professional Maintenance - Chapter - 3 - ADocument100 pagesWCM - Yamashina - 05 - Professional Maintenance - Chapter - 3 - ACíceroNo ratings yet
- Rice 1788Document70 pagesRice 1788Dr. Rohit SinghNo ratings yet
- Jurnal Fatimah SimbolonDocument8 pagesJurnal Fatimah SimbolonFatimah SimbolonNo ratings yet
- tp2 Systc3a8mes Asservis Numc3a9riquesDocument1 pagetp2 Systc3a8mes Asservis Numc3a9riquesomarNo ratings yet
- CHT307 - Ktu QbankDocument3 pagesCHT307 - Ktu QbankYuxin CasioNo ratings yet
- CBSE Class 12 Chemistry Quick Revision Notes Co-Ordination CompoundsDocument8 pagesCBSE Class 12 Chemistry Quick Revision Notes Co-Ordination CompoundsAbid waniNo ratings yet
- Ip Rating PDFDocument1 pageIp Rating PDFGurunadha Rao RamachandraNo ratings yet
- Isolated Footing DesignDocument106 pagesIsolated Footing DesignAlemayehu Miteku100% (1)
- Microscope Laboratory Activities 2023Document7 pagesMicroscope Laboratory Activities 2023Izzy Dale Cagas SilangaNo ratings yet
- Module 4 Work Energy PowerDocument7 pagesModule 4 Work Energy PowerGreen BrainNo ratings yet
- Gujarat Technological UniversityDocument1 pageGujarat Technological UniversityBhagirath sinh ZalaNo ratings yet
- Fujifilm Eluxeo BL 7000Document78 pagesFujifilm Eluxeo BL 7000AngeloNo ratings yet
- RadioaktivlikDocument3 pagesRadioaktivlikgultaczaurNo ratings yet
- Synthesis of Zns / Sodium Hexameta Phosphate Nanoparticles: R. Mohan, B. Rajamannan and S. SankarrajanDocument6 pagesSynthesis of Zns / Sodium Hexameta Phosphate Nanoparticles: R. Mohan, B. Rajamannan and S. SankarrajanphysicsjournalNo ratings yet
- Current Logic 1000V - 24V Power SupplyDocument2 pagesCurrent Logic 1000V - 24V Power Supplyraza239No ratings yet
- AC Induction Motor MC WebinarDocument61 pagesAC Induction Motor MC WebinarAdeNo ratings yet
- Cape Wind 462 MW GenerationDocument78 pagesCape Wind 462 MW GenerationmilanovsdNo ratings yet
- EE6512-Electrical Machines LaboratoryDocument82 pagesEE6512-Electrical Machines LaboratoryGopinath B L NaiduNo ratings yet
- s4 Physics Paper 3 Exam 5Document4 pagess4 Physics Paper 3 Exam 5MUSOKE GERALDNo ratings yet
- Math Reviewer Lesson 1 3Document27 pagesMath Reviewer Lesson 1 3Junel BitangaNo ratings yet
- Pamantasan NG Lungsod NG ValenzuelaDocument2 pagesPamantasan NG Lungsod NG ValenzuelaDiane de OcampoNo ratings yet
- Mechanics of Fluids Si Edition 5th Edition Potter Test BankDocument3 pagesMechanics of Fluids Si Edition 5th Edition Potter Test Bankwalleyeemploye2g100% (28)
- ENE - Combined Updated For Use of AY2020Document2 pagesENE - Combined Updated For Use of AY2020Alzheimer RachNo ratings yet
- 425ºC AHT: Ower EATDocument2 pages425ºC AHT: Ower EATSivagurunathan SpNo ratings yet
(W-6302) Neural Mode Jump Monte Carlo
(W-6302) Neural Mode Jump Monte Carlo
Uploaded by
Nazmul SaqibOriginal Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
(W-6302) Neural Mode Jump Monte Carlo
(W-6302) Neural Mode Jump Monte Carlo
Uploaded by
Nazmul SaqibCopyright:
Available Formats
Neural mode jump Monte Carlo
Accepted: 1 January 2021 •
Luigi Sbailò,1 Manuel Dibak,1 and Frank Noé1,2,a)
ABSTRACT
Markov chain Monte Carlo methods are a powerful tool for sampling equilibrium configurations in complex systems. One problem these
methods often face is slow convergence over large energy barriers. In this work, we propose a novel method that increases convergence in
systems composed of many metastable states. This method aims to connect metastable regions directly using generative neural networks
in order to propose new configurations in the Markov chain and optimizes the acceptance probability of large jumps between modes in
the configuration space. We provide a comprehensive theory as well as a training scheme for the network and demonstrate the method on
example systems.
02 February 2024 01:29:53
I. INTRODUCTION temperature of the sampler from a set of discrete temperatures
while remaining at equilibrium in an augmented temperature-
Markov chain Monte Carlo (MCMC) methods are used to sam- configuration space. In parallel tempering, multiple simulations at
ple the equilibrium distribution of systems whose probability dis- different temperatures are carried out in parallel and samples are
tribution is otherwise analytically intractable. An efficient MCMC randomly exchanged between the different temperatures. These
generator proposes moves that quickly decorrelate the samples while methods rely on a significant overlap of the energetic distributions
having a large acceptance probability. The Metropolis–Hastings at different temperatures; therefore, the temperature range has to be
algorithm1 is a standard method, where trial moves are generated chosen carefully. Even though it is possible to optimize the num-
with a random displacement in the configuration space. As com- ber of replicas using nonequilibrium simulations,8 the efficiency of
plex systems at equilibrium visit only a small fraction of the whole the scheme still depends on the chosen dynamics. In the popula-
configuration space, these random displacements have to be very tion annealing algorithm,9 many replicas of the system are initial-
small to be accepted. However, small moves are only efficient at ized at high temperatures and then the population is cooled down.
sampling local conformations of the energy landscape, while cross- A resampling step ensures that replicas are kept near equilibrium,
ing large energy barriers requires a multitude of sampling steps. but statistical correlations between replicas might lead to systematic
This problem is particularly evident when the system is composed errors.10
of many metastable states,2 where it is often computationally infea- Several methods utilize a pre-defined reaction coordinate
sible to cross energy barriers multiple times, and convergence to between metastable states to enhance sampling. A class of meth-
equilibrium would require an enormous amount of computational ods biases the potential landscape along the reaction coordinate to
time. enable transitions over energy barriers and recovers the unbiased
In the last few decades, many different methods have been distribution by re-weighting. Metadynamics11 does this in an iter-
developed to circumvent this problem: One class of methods varies ative fashion, where the bias potential is increased in areas where the
the temperature during the sampling process as the crossing time system resides a long time, thus pushing the system out of metastable
over energy barriers exponentially decreases with inverse temper- states. Recent developments suggest the usage of deep learning to
atures. The two most widely recognized methods in this class are find an optimal bias potential.12 Umbrella sampling13 runs several
simulated3,4 and parallel tempering,5–7 which operate on a fixed sampling iterations with bias potentials placed along the reaction
set of temperatures. Simulated tempering randomly changes the coordinate and thus pushes the system from one end to the other.
J. Chem. Phys. 154, 074101 (2021); doi: 10.1063/5.0032346 154, 074101-1
© Author(s) 2021
The Journal
ARTICLE scitation.org/journal/jcp
of Chemical Physics
Nonequilibrium path sampling methods14–16 construct which could be obtained from, e.g., x-ray scattering or nuclear mag-
reversible moves between equilibrium states as a collection of small netic resonance (NMR) experiments from which starting points in
out-of-equilibrium trajectories. These moves are typically generated different conformations can be generated.31 Possible applications
by driving the system along the reaction coordinate, and a system range from proteins with multiple metastable states (e.g., Kinases)
specific protocol to generate the proposed state must be designed. to solid state systems with multiple phases, where configurations in
A recent approach uses machine learning methods in order to learn either phase can be easily generated, but observing the transition
such a protocol.17 is rare. Local displacements and neural moves are randomly alter-
In situations where states are disconnected and finding a reac- nated in a combined scheme to accelerate the convergence rate of
tion coordinate is challenging, different approaches have been devel- Markov chains. Configurations from different metastable states are
oped. Smart darting Monte Carlo18,19 is a promising method that used to train the networks, which are optimized to produce high
alternates local and long range moves from one region of the con- acceptance probability moves. Local exploration ensures ergodicity
figuration space to another that is arbitrary far. These moves are of the scheme, while neural moves accelerate convergence to equilib-
attempted between small spheres around local minima. In high rium, realizing an accurate and deep exploration of the configuration
dimensions, however, the fraction of the spheres in the total volume space.
becomes vanishingly small, and therefore, finding a sphere by ran-
dom exploration becomes unlikely. This problem is circumvented in
ConfJump20 by finding the closest energy minimum and attempting II. THEORY
long range moves by translation to another energy minimum. A sufficient condition to ensure that a Markov chain asymptot-
The generation of long range moves is challenging when the ically samples the equilibrium distribution is ergodicity and detailed
energy landscape is rough since the potential energy surface in the balance.32 Given the system in a configuration x, a new state y is
region surrounding local minima can drastically change among the added to the chain according to a transition probability p(x → y).
different minima. In this case, using trivial translation as long range The transition probability is defined to satisfy the condition of
moves would most likely cause large energy differences, and trial detailed balance
moves are likely to be rejected. In fact, in order to keep the energy
difference small, trial moves should be generated with a bijective π(x) p(x → y) = π(y) p(y → x), (1)
function that pairs points with similar energy values. Constructing
such bijection manually would require detailed knowledge of the
02 February 2024 01:29:53
system and is practically impossible in multi-dimensional systems. where π(x) is the stationary distribution.33 In the Metropolis–
On the other hand, recent advances in the field of machine learning Hastings algorithm,1,32 the transition probability is decomposed into
have permitted to deal with problems that were not solvable with a two logical steps: first, a new configuration y is drawn from a pro-
sole human understanding, and more specifically, deep neural net- posal density pprop (x → y), and then, the new state is accepted with
works (DNNs) are an ideal tool to facilitate the approximation of a an acceptance probability pacc (x → y). If the transition is accepted,
desired function. the new state y is added to the Markov chain; otherwise, the previous
DNNs have already been employed to construct MCMC moves. state x is added to the Markov chain.
Current methods use DNNs for approximating the target distribu- In Neural MJMC, the proposal probability is split into two
tion21 or projecting onto high probability manifolds.22 Two recent steps: first, a proposal density is selected from a pre-defined list of
methods use reversible network architectures to improve Hamilto- proposal densities on the current state x, and then, a new state y is
nian Monte Carlo (HMC): A-NICE-MC23 proposes steps by apply- drawn from the extracted proposal density. Proposal densities are
ing a volume preserving flow to the augmented configuration space. distinguished between local proposals and neural proposals, where
Another method24 augments the leapfrog algorithm commonly used local proposals generate local moves, e.g., through random dis-
in HMC with DNNs and thus alters the classical path of the system placement with Gaussian increments, and neural proposals connect
while relying on forces. Both are trained for sampling efficiency in different metastable states with global moves.
an unsupervised fashion and therefore rely on random exploration Let us assume that the configuration space Ω is decomposed
of the configuration space in order to find metastable states. into a number of non-overlapping subsets called cores {Ωα }α≤N
In Boltzmann generators,25,26 reversible DNNs are used to draw ⊂ Ω, with ∪α Ωα = Ω, each representing one of the N metastable
statistically independent equilibrium samples of condensed matter states. We define the neural proposal K αβ as the density that pro-
systems and protein molecules in “one shot.” This method uses poses transitions from the core Ωα to the core Ωβ . Assuming that
deep learning for sampling a target distribution by combining an the system is in state x ∈ Ωα , the probability to extract the neural
exact probability generator such as a normalizing flow27,28 with proposal K αβ is pαβ (x). Once K αβ has been selected, a state y ∈ Ωβ is
αβ
reweighting.29,30 drawn from the selection probability pprop (x → y).
In this paper, we present neural mode jump Monte Carlo (Neu- A neural proposal K αβ can only be selected within the core
ral MJMC), a novel method to efficiently sample the equilibrium Ωα and with constant probability pαβ (x) = pαβ χΩα (x), where χ Ω (x)
distribution of complex many-body systems with unbiased Markov denotes the characteristic function. We assume that each pair of
chains. In this scheme, reversible neural networks are trained to states (α, β) is only connected by one neural proposal K αβ and that
propose “neural” moves that directly connect different metastable there exists an inverse proposal K βα connecting β with α. Under
states as neural moves do not depend on the physical path connect- these assumptions, a proposed move starting in Ωα with selected
ing the start and endpoint. The method requires a prior knowledge neural proposal K αβ fulfills detailed balance if it is accepted with
of the position of the metastable states in the configuration space, probability
J. Chem. Phys. 154, 074101 (2021); doi: 10.1063/5.0032346 154, 074101-2
© Author(s) 2021
The Journal
ARTICLE scitation.org/journal/jcp
of Chemical Physics
⎧
⎪ ⎫
⎪ π(y) pβα pprop (y → x) ⎪
βα
αβ ⎪ ALGORITHM 1. Neural MJMC sampling scheme.
pacc (x → y) = min⎨1, ⎬. (2)
⎪
⎪ ⎪
⎩ π(x) pαβ pprop (x → y) ⎪
αβ
⎭ input: ls = []: empty list for samples
{pα β} : proposal selection probabilities
We parameterize the neural proposal K αβ and its inverse K βα con-
{μα β}: proposal densities
necting the cores Ωα and Ωβ as a bijective function μαβ (⋅) pairing the
x ← x0 : starting point of sampling
states defined in the two cores, i.e., y = μαβ (x), μ−1
αβ (y) = x, ∀x ∈ Ωα , N iterations : number of generated samples
where x ∈ Ωα , y ∈ Ωβ . Thus, for each pair of different cores (Ωα , Ωβ ), σ local : standard deviation of local moves
a bijective function μαβ (⋅) is defined. The probability distribution of while i ≤ N iterations do
neural proposals is then represented with Dirac delta distributions draw proposal density K αβ from {pαβ }
and the acceptance specifies to /if α = β then //propose local move
w ← sample from N(0, 𝟙)
⎧
⎪ −1 ⎫ / y ← x + w ⋅ σ local
αβ ⎪ π(y) pαβ δ(x − μαβ (y)) ⎪ ⎪ /
pacc (x → y) = min⎨1, ⎬. (3)
⎪
⎪ π(x) pβα δ(y − μαβ (x)) ⎪
⎪ pacc ← pαα acc (x → y) [Eq. (5)]
⎩ ⎭ /else //propose neural move
y = μαβ (x)
Using the change of variable formula in the Dirac distribution / /p ← pαβ (x → y) [Eq. (4)]
δ(x − μ−1
αβ (y)) = ∣det Jμαβ (x)∣δ(y − μαβ (x)) with the Jacobian Jμαβ (x)
acc acc
of the function μαβ , the acceptance probability for neural moves can if r ∼ U(0, 1) < pacc then
/ /x←y
be simplified to
end
/ ls .append (x)
αβ π(y) pαβ
pacc (x → y) = min{1, ∣det Jμαβ (x)∣}. (4) / i←i+1
π(x) pβα
end
output: list of samples ls
In case the local proposal (α = β) is selected, the inverse move is only
possible with another local proposal K αα . Note that a local move may
leave the current core and the proposal probability for the inverse
02 February 2024 01:29:53
move might change. Thus, the acceptance probability for a local
move reduces to min E[β∣ΔVαβ (x) + kB T log∣det Jμαβ (x)∣ + ΔRαβ ∣], (7)
μαβ
π(y) ∑β χΩβ pββ
pαα
acc (x → y) = min{1, }. (5) with the potential difference ΔV αβ (x) ∶= V(x) − V(μαβ (x)) and the
π(x)pαα log selection ratio ΔRαβ ∶= − kB T log pαβ /pβα . Note that the term
inside the modulus is equivalent to the Kullback–Leibler diver-
In order to ensure ergodicity, there needs to be a finite probabil- gence between the transformed distribution μαβ (Ωα ) and the target
ity of selecting the local proposal in all cores. In Algorithm 1, we distribution Ωβ as found in Ref. 26.
summarize the Neural MJMC sampling scheme. We can interpret this result in a physically meaningful man-
ner by applying the triangular inequality E[−∣log f ∣] ≥ −∣E[log f ]∣,
A. Optimal proposal density identifying ΔS = −kB E[log∣det Jμαβ (x)∣] as the change of differen-
To achieve fast decorrelation of the Markov chain, the neu- tial entropy (see Appendix A for details) and ΔU = E[ΔVαβ (x)] as
ral proposal functions μαβ should maximize the acceptance in both the change of internal energy under the transformation μαβ (x). We
directions. This is quantified by maximizing the expected log proba- observe that the expected log acceptance is lower bound by the abso-
bility that the moves proposed by μαβ are accepted in both directions. lute change in free energy ΔF = ΔU − TΔS under the transformation
Using Jensen’s inequality, we find μij (⋅) divided by thermal energy
max log Ex∼Ωα [ ppacc (x → y)ppacc (y → x)] E{log[ ppacc (x → y)ppacc (y → x)]} ≥ −β∣ΔF + ΔRij ∣. (8)
μαβ
≥ max E{log[ ppacc (x → y)ppacc (y → x)]} This result shows that we can use the freedom in the proposal
= max E[min(0, log f ) + min(0, − log f )] selection ratio to maximize the bi-directional acceptance.
μαβ
= max E[min(log f , − log f )] = max E[−∣log f ∣], (6)
μαβ μαβ III. NEURAL NETWORK ARCHITECTURE
π(y) p As neural moves are defined with invertible functions, it is ben-
where f = π(x) pαβ
βα
∣det J(μαβ (x))∣. Using the stationary distribution eficial to choose neural networks that are invertible. Recently, several
in the canonical ensemble π(x) ∝ exp(−βV(x)), with the thermal invertible architectures have been proposed23,27,28,34 with variable
energy β−1 = kB T and the potential energy V(x) of the system under expressiveness and computational cost.
consideration, and assuming that μαβ is a bijection between the cores In order to ensure that the output of the network μαβ (⋅) is in the
(α, β), we can rewrite the above equation to find correct well, a harmonic bias potential centered in the target core is
J. Chem. Phys. 154, 074101 (2021); doi: 10.1063/5.0032346 154, 074101-3
© Author(s) 2021
The Journal
ARTICLE scitation.org/journal/jcp
of Chemical Physics
added during training,
⎧
⎪
⎪k(x − xα ) x ∈ Ωα
2
Vbias (x) = ⎨ (9)
⎪ 2
⎩k(x − xβ )
⎪ x ∈ Ωβ ,
where xα is the reference configuration in core α and the spring con-
stant k is a hyperparameter, resulting in the biased system Ṽ(x)
= V(x) + Vbias (x) used during training. The network is trained in
several stages, gradually lowering the strength of the bias potential.
To find the reference configurations xα , k-means clustering is run
on samples generated from local MCMC sampling in either well.
Training sets of both of the wells are generated for a set of grad-
ually decreasing bias strengths {ki }i≤Nk . After convergence of the
training at ki , the training set is exchanged and training is restarted
with ki+1 ≤ ki . This allows for a slowly expanding training set, which
enables the network to learn how to generate meaningful moves
on a gradually more complex set of training data. The loss that
is to be minimized during training is given by the bi-directional
acceptance (7)
2
Cacc = Ex∼Ωα {[ΔṼαβ (x) + kB T log∣det Jμαβ (x)∣] }, (10)
where the square of the norm is used to penalize high energies.
Training is performed in the forward and backward directions, and
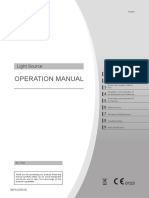
the same loss applies to samples from core Ωβ with exchanged labels FIG. 1. Top: schematic figure of the Neural MJMC scheme. Given configuration
α ↔ β. x in core Ωα , there are three neural proposals and one local proposal available,
02 February 2024 01:29:53
as denoted by arrows. One of these is selected, and a new state y is proposed.
Bottom: architecture of the RNVP networks that are used as reversible networks for
IV. NUMERICAL EXPERIMENTS the examples in this paper. The input configuration x is separated into two disjoint
(0) (0) (0)
We demonstrate Neural MJMC on two examples: a two- sets of coordinates x1 , x2 . At the first iteration, only subsets x2 are fed into
dimensional potential landscape with three minima and a system the neural networks S0 , T 0 allowing for nonlinear transformations. The outputs of
the neural networks S0 , T 0 are, respectively, multiplied and added to the other
consisting of two dimer particles that are suspended in a bath (0) (0)′ (0)
of repulsive particles. The functional form of the potentials and subset x1 to arrive at x1 , while x2 is kept constant. In the following iteration,
(0)′
detailed training parameters are given in Appendixes B and C. The the subsets are swapped and x2 gets transformed by functions depending only
(0)′
code required to generate the results is available on github. on x1 . Stacking many of these iterations allows for a complex transformation
As a good compromise between computational cost and that can be easily inverted.
expressiveness, we use real non-volume preserving (RNVP) trans-
formations27 for these examples. In a RNVP layer, the configuration
tensor x ∈ RN×dim is split into two vectors x1 and x2 . As we deal with
two-dimensional systems, we split along the x and y coordinates of
A. Gaussian triple well
all particles such that x1 , x2 ∈ RN×1 . One RNVP layer consists of two
update steps in which the first subset is updated based on the second, As an example for a system with multiple states, we demon-
while the second is kept constant and vice versa, strate Neural MJMC on a two-dimensional potential landscape con-
sisting of three Gaussian shaped wells. We define the three cores by a
⎡x(i)′⎤ ⎡ (i) (i) (i) ⎤
⎢1 ⎥ ⎢ x1 ⊙ exp[Si (x2 )] + Ti (x2 )⎥ Voronoi tessellation36 for which we use the minima of the Gaussians
⎢ (i)′⎥ = ⎢ ⎥,
⎢x ⎥ ⎢ ⎥
(i)
(11) as centers. Each neural network is trained independently on config-
⎣2 ⎦ ⎢ ⎣ x2 ⎥
⎦ urations sampled from the minima. In the sampling step, 100 inde-
pendent trajectories of length 105 steps are generated and averaged.
(i+1) ⎡ (i)′ ⎤
x ⎢ x1 ⎥ We compare the marginal distributions pX and pY , which are the
[ 1(i+1)] = ⎢ ′ ′ ′ ⎥,
⎢x(i) ⊙ exp[S′ (x(i) )] + T ′ (x(i) )⎥ (12) projections of the Boltzmann distribution on the X and Y axes, and
x2 ⎢2 ⎥
⎣ i 1 i 1 ⎦ observe great agreement to results from the numerical integration of
the Boltzmann distribution (see Fig. 2).
where the Si , T i , T i′ , Si′ are dense feedforward neural networks.35
The above system of equations represent one RNVP block, and arbi-
B. Dimer in repulsive Lennard-Jones bath
trary number of these blocks can be serially stacked, resulting in
more complex transformations (see Fig. 1). The logarithm of the As a bigger challenge, Neural MJMC is applied to a two-
Jacobian determinant of this transformation is given by the sum over dimensional system composed of a bistable dimer immersed in
all outputs of all the scaling layers log∣det Jμαβ ∣ = ∑i (Si + S′i ). a bath of strongly repelling particles and confined to a box. The
J. Chem. Phys. 154, 074101 (2021); doi: 10.1063/5.0032346 154, 074101-4
© Author(s) 2021
The Journal
ARTICLE scitation.org/journal/jcp
of Chemical Physics
FIG. 2. Two-dimensional histogram (center) of samples from the 2D Gaussian
triple well potential generated by Neural MJMC with a short section of the Markov
chain (yellow solid line) and marginal distributions pX (top) and pY (right). The
black dashed line depicts the border between the states that are defined by a
Voronoi tessellation. Convergence to the correct Boltzmann distribution can be
observed from the histograms of the marginal distributions, where the red line
02 February 2024 01:29:53
is the reference solution from numerical integration of the system’s Boltzmann
distribution.
FIG. 3. Top: free energy along the distance between the dimer particles. The cor-
responding bands represent reference values obtained by umbrella sampling with
bistable dimer potential has a minimum in the closed and open the standard error given by their thickness. The neural network has been trained
configurations, which are separated by a high energy barrier (see at temperature T = T 0 ; then, simulations at different temperatures have been per-
Fig. 3, middle). Opening and closing of the dimer require a concerted formed using Neural MJMC. Simulations are run for 1.5 × 107 steps, and error
motion of the solvent particles, which makes it difficult to sample the bars are generated from several sampling runs. In this figure, we observe that
physical path connecting the two configurations (see Appendix B for Neural MJMC correctly samples the free energy along the reaction coordinate of
the system at different temperatures. Middle: dimer interaction potential E d as a
a more detailed description of the system). function of the dimer distance. Bottom: reference configurations in the closed (left)
The open and closed configurations serve as cores (see Fig. 3, and open (right) dimer configuration. The dimer particles are displayed in blue,
bottom) in Neural MJMC and are distinguished by the distance and solvent particles are displayed in gray. The strongly repulsive potential does
between the dimer particles. The neural network is trained on states not allow for significant overlaps between particles at equilibrium.
sampled independently in the closed and open configurations at four
different bias strengths with 105 samples for each well and bias. As
the system is invariant under permutation of solvent and dimer par-
ticles, neural moves would have to be learned independently for each for this system. This slowdown arises from the evaluation of the net-
permutation of the system, which is clearly unfeasible as the number work and the remapping of particles. As a reference value, we use
of permutation scales factorially with the number of particles. This umbrella sampling13 to sample the free energy along the dimer dis-
problem is circumvented by permutation reduction, i.e., relabeling tance. To this end, we use 20 umbrellas along the dimer distance and
the particles such that the distance to the reference configuration is compute configurations with MCMC. The free energy is calculated
minimized, which is realized using the Hungarian algorithm37 with using the Multistate Bennet Acceptance Ratio38 method.
the reference configurations as the target. Neural moves cause direct transitions between the two
Each neural network in the RNVP architecture consists of three metastable states and thus a rapid exploration of the configuration
hidden layers with 76 nodes. The transformation consists of a total of space. The convergence to the Boltzmann distribution is observed as
20 RNVP layers and contains ∼1.4 × 106 trainable parameters. Neu- shown in Fig. 3. An estimate for the crossing time with only local
ral MJMC is used to generate a single trajectory with 1.5 × 107 steps, moves can be found to be at the order of 1012 sampling steps at T = 1
where the probability of neural moves is set to 1%. In terms of com- from the Kramers problem,39 which makes exhausting simulations
putational performance, sampling with Neural MJMC is approxi- unfeasible. In Neural MJMC, many crossings of the energy barrier
mately a factor of four slower than MCMC with local displacements can be observed (Fig. 4, top). This is also reflected in the autocorre-
J. Chem. Phys. 154, 074101 (2021); doi: 10.1063/5.0032346 154, 074101-5
© Author(s) 2021
The Journal
ARTICLE scitation.org/journal/jcp
of Chemical Physics
Reversible neural network architectures are also used in the
field of generative probabilistic modeling and have been success-
fully applied in machine learning frameworks for high-dimensional
problems such as generation of high-resolution images, graphs,
and molecular structures. Considering the great attention this field
is lately facing, it would not be surprising to observe dramatic
improvements in the performance of such reversible networks. An
important step to further generalization is the incorporation of
physical symmetries into the network architecture. Many recent
advances when applying machine learning to physical problems
make use of a symmetric representation.40–42 This has also been
applied to the field of invertible networks,43 and using an architec-
ture that preserves symmetries appears promising to allow for scal-
ing of the proposed method to even bigger systems. For molecules,
such a representation can be found in terms of bonds and angles.
This has recently been shown to be able to reproduce the distri-
bution of configurations for smaller molecules.17 Such an approach
could straightforwardly be applied in this method in future work.
The efficiency and capability of neural MJMC profoundly rely on
the specific architecture employed, and more sophisticated networks
would allow us to deal with systems of increasing complexity. Neural
MJMC is a general and transferable method, and we can expect it to
be applied to a multitude of different systems.
FIG. 4. Top: dimer distance over a single realization using Neural MJMC (top)
and using local MCMC (bottom). (Right) Histogram of the dimer distance obtained
by the displayed trajectory, with the reference value displayed as the black dashed AUTHORS’ CONTRIBUTIONS
line. Spontaneous transitions with local MCMC are not observed at this time-scale.
02 February 2024 01:29:53
Neural MJMC explores both metastable states in the trajectory multiple times and L.S. and M.D. authors contributed equally to this work.
correctly reproduces the distribution of dimer distances. Bottom: autocorrelation of
the dimer distance. Neural moves allow for a fast exploration of both metastable
states, accelerating the production of uncorrelated samples. In this figure, it is ACKNOWLEDGMENTS
evident that Neural MJMC frequently generates uncorrelated samples, and short
trajectories are sufficient to reconstruct the right distribution. In contrast, configu- We gratefully acknowledge funding from the European Com-
rations generated with local MCMC are highly correlated as they do not cross the mission (Grant No. ERC CoG 772230 “ScaleCell”), Deutsche
energy barrier. Forschungsgemeinschaft (Grant No. CRC1114/C03), the
MATH+ Berlin Mathematics Research Center (Grant No. AA1-
6), and the International Max Planck Research School IMPRS-
CBSC. Furthermore, we want to thank Christoph Fröhner, Mohsen
Sadeghi, and Andreas Mardt for insightful discussions.
lation function where samples generated with local MCMC remain
highly correlated, while it decays in Neural MJMC simulations on a
scale of ∼105 sampling steps (see Fig. 4, bottom), thus generating the APPENDIX A: DIFFERENTIAL ENTROPY DIFFERENCE
desired uncorrelated samples of the equilibrium distribution. The differential entropy of a system is defined as
V. CONCLUSION SX = −kB ∫ pX (x) log pX (x)dx. (A1)
Ω
In this paper, we have presented neural mode jump Monte
Carlo, a novel method that allows for efficient sampling of the For a bijective function y = μαβ (x), we can apply the change
Boltzmann distribution of complex systems composed of many of variable formula to compute the change in differential entropy
metastable states. The method uses neural networks in order to under the transformation. With the transformed density being
parameterize bijections between metastable regions in phase space pY (y) = pX (μ−1
αβ (y))∣det Jμ−1 (y)∣, we find
αβ
and optimizes these networks for bi-directional acceptance proba-
bility. By combining short steps given by random displacements and
large jumps between metastable states, the method is able to con- SX = −kB ∫ pY (y) log pY (y)∣det Jμ−1 (y)∣dy
μαβ (Ω) αβ
verge quickly to the Boltzmann distribution. This is especially evi-
dent in systems where large potential barriers are providing obstacles = SY − kB ∫ pY (y) log∣det Jμ−1 (y)∣dy. (A2)
to the convergence of other methods. The method is demonstrated μαβ (Ω) αβ
on two toy examples, one with several bijections in two dimensions
and the other with a high-dimensional system consisting of a particle Thus, the difference in differential entropy under the transfor-
dimer in a bath of Lennard-Jones particles. mation μαβ (⋅) is given as
J. Chem. Phys. 154, 074101 (2021); doi: 10.1063/5.0032346 154, 074101-6
© Author(s) 2021
The Journal
ARTICLE scitation.org/journal/jcp
of Chemical Physics
ΔS = SY − SX = −kB Ex∼pΩ [log∣det Jμαβ (x)∣], (A3) TABLE III. Parameters of the RNVP networks used in the experiments.
where we used the inverse function theorem to compute the DW Particles
Jacobian.
Number of blocks 10 20
Hidden dimensions [20, 20, 20] [76, 76, 76]
APPENDIX B: FUNCTIONAL FORM OF POTENTIALS Number of parameters 3.6 × 104 1.4 × 106
Here, we give the exact functional form of the potentials used Number of training samples 1 × 10 5
1 × 105
to demonstrate the proposed method. per bias and core
Bias strengths/kB T [10, 0] [500, 10, 5, 2]
1. Triple well potential Learning rate 10–3 [10− 3 , 10− 4 , 10− 4 , 10− 5 ]
Batch size 2000 8192
The triple well potential is a 2D potential surface given by
V(x) = ∑ −ai exp[−(x − mi )T Σi (x − mi )] + b∥x∥2 , (B1)
i
APPENDIX C: DETAILS OF THE NETWORK
with b = 0.1 and other parameters given in Table I. ARCHITECTURE
The RNVP network consists of several subsequent blocks as
2. Dimer in a Lennard-Jones bath depicted in Fig. 1 (bottom). Each of these blocks consists of four
The dimer system is adapted from Ref. 26. It consists of N = 38 independent networks, two for scaling and two for translation. All
particles in two dimensions. Its potential energy is given by networks use leaky ReLU in each hidden layer. The output of the
scaling networks uses a hyperbolic tangent scaled by a trainable
U(x1 , . . . , xN ) = Ubox (x1 , . . . , xN ) + Urep (x1 , . . . , xN ) scalar. The output of the translation networks is linear. Adam44
+ Ud (x1 , x2 ), is used as the optimizer with standard parameters and a learn-
ing rate depending on the system. Table III gives an overview of
02 February 2024 01:29:53
with the exact network architectures and hyperparameters used in the
N 2 experiments.
Ubox (x1 , . . . , xN ) = kbox ∑ ∑ Θ(∣xi,d ∣ − lbox )2 ,
i=1 d=1
DATA AVAILABILITY
−1 −12
Urep (x1 , . . . , xN ) = ϵ ∑ ∑ (rm ∥xi − xj ∥2 ) , The data that support the findings of this study are available
j≤N i<j
(i,j)≠(1,2) from the corresponding author upon reasonable request. The com-
puter code required to generate the results presented in this paper is
Ud (x, y) = ad(x, y) + bd(x, y)2 + cd(x, y)4 available here.
+ kd [(x1 + y1 )2 + x22 + y22 ],
REFERENCES
with d(x, y) = 2(∥x − y∥22 − d0 ) and the Heaviside step function 1
N. Metropolis, A. W. Rosenbluth, M. N. Rosenbluth, A. H. Teller, and E. Teller,
Θ(x). The specific parameters used in this paper are given in “Equation of state calculations by fast computing machines,” J. Chem. Phys. 21,
Table II. 1087–1092 (1953).
2
H. Frauenfelder, S. G. Sligar, and P. G. Wolynes, “The energy landscapes and
motions of proteins,” Science 254(5038), 1598–1603 (1991).
3
TABLE I. Parameters of the triple well potential. C. J. Geyer and E. A. Thompson, “Annealing Markov chain Monte Carlo with
applications to ancestral inference,” J. Am. Stat. Assoc. 90(431), 909–920 (1995).
4
i Σi mTi ai E. Marinari and G. Parisi, “Simulated tempering: A new Monte Carlo scheme,”
Europhys. Lett. 19(6), 451–458 (1992).
5
1 diag(0.5, 0.3) (−2.2, −1) 5 C. J. Geyer, “Markov chain Monte Carlo maximum likelihood,” in Computing
Science and Statistics: Proceedings of the 23rd Symposiumon on the Interface
2 diag(0.5, 0.4) (0, 2) 5 (Interface Foundation of North America, 1991).
3 diag(0.4, 0.5) (2, −0.8) 5 6
K. Hukushima and K. Nemoto, “Exchange Monte Carlo method and application
to spin glass simulations,” J. Phys. Soc. Jpn. 65(6), 1604–1608 (1996).
7
R. H. Swendsen and J.-S. Wang, “Replica Monte Carlo simulation of spin-
glasses,” Phys. Rev. Lett. 57, 2607–2609 (1986).
TABLE II. Parameters of the particle dimer system. 8
A. J. Ballard and C. Jarzynski, “Replica exchange with nonequilibrium switches,”
Proc. Natl. Acad. Sci. U. S. A. 106(30), 12224–12229 (2009).
9
Parameter kbox N lbox ϵ rm d0 a b c kd K. Hukushima and Y. Iba, “Population annealing and its application to a spin
glass,” AIP Conf. Proc. 690(1), 200–206 (2003).
Value 100 36 3.0 1.0 1.0 1.5 0 −25 10 20 10
C. Amey and J. Machta, “Analysis and optimization of population annealing,”
Phys. Rev. E 97, 033301 (2018).
J. Chem. Phys. 154, 074101 (2021); doi: 10.1063/5.0032346 154, 074101-7
© Author(s) 2021
The Journal
ARTICLE scitation.org/journal/jcp
of Chemical Physics
11 29
A. Laio and M. Parrinello, “Escaping free-energy minima,” Proc. Natl. Acad. Sci. M. S. Albergo, G. Kanwar, and P. E. Shanahan, “Flow-based generative models
U. S. A. 99(20), 12562–12566 (2002). for Markov chain Monte Carlo in lattice field theory,” Phys. Rev. D 100, 034515
12 (2019).
J. Zhang, Y. I. Yang, and F. Noé, “Targeted adversarial learning optimized
30
sampling,” J. Phys. Chem. Lett. 10(19), 5791–5797 (2019). K. A. Nicoli, S. Nakajima, N. Strodthoff, W. Samek, K.-R. Müller, and P. Kessel,
13 “Asymptotically unbiased generative neural sampling,” Phys. Rev. E 101(2),
G. M. Torrie and J. P. Valleau, “Nonphysical sampling distributions in Monte
Carlo free-energy estimation: Umbrella sampling,” J. Comput. Phys. 23(2), 187– 023304 (2020).
31
199 (1977). D. L. Parton, P. B. Grinaway, S. M. Hanson, K. A. Beauchamp, and J. D.
14
Y. Chen and B. Roux, “Constant-pH hybrid nonequilibrium molecular dynam- Chodera, “Ensembler: Enabling high-throughput molecular simulations at the
ics Monte Carlo simulation method,” J. Chem. Theory Comput. 11(8), 3919–3931 superfamily scale,” PLoS Comput. Biol. 12(6), e1004728 (2016).
32
(2015). W. K. Hastings, “Monte Carlo sampling methods using Markov chains and their
15
J. P. Nilmeier, G. E. Crooks, D. D. L. Minh, and J. D. Chodera, “Nonequilibrium applications,” Biometrika 57(1), 97–109 (1970).
candidate Monte Carlo is an efficient tool for equilibrium simulation,” Proc. Natl. 33
C. W. Gardiner, Handbook of Stochastic Methods for Physics, Chemistry and the
Acad. Sci. U. S. A. 108(45), E1009–E1018 (2011). Natural Sciences, Springer Series in Synergetics, 3rd ed. (Springer-Verlag, Berlin,
16
H. A. Stern, “Molecular simulation with variable protonation states at constant 2004), Vol. 13.
pH,” J. Chem. Phys. 126(16), 164112 (2007). 34
T. Q. Chen, Y. Rubanova, J. Bettencourt, and D. K. Duvenaud, “Neural ordi-
17
H. Wu, J. Köhler, and F. Noé, “Stochastic normalizing flows,” arXiv:2002.06707 nary differential equations,” in Advances in Neural Information Processing Systems
(2020). (Curran Associates, Inc., 2018), pp. 6571–6583.
18 35
I. Andricioaei, J. E. Straub, and A. F. Voter, “Smart darting Monte Carlo,” P. Mehta, M. Bukov, C.-H. Wang, A. G. R. Day, C. Richardson, C. K. Fisher,
J. Chem. Phys. 114(16), 6994–7000 (2001). and D. J. Schwab, “A high-bias, low-variance introduction to machine learning
19
K. Roberts, R. Sebsebie, and E. Curotto, “A rare event sampling method for for physicists,” Phys. Rep. 810, 1–124 (2019), part of the Special Issue: A high-bias,
diffusion Monte Carlo using smart darting,” J. Chem. Phys. 136(7), 074104 low-variance introduction to Machine Learning for physicists.
36
(2012). G. Voronoi, “Nouvelles applications des paramètres continus à la théorie
20
L. Walter and M. Weber, “ConfJump: A fast biomolecular sampling method des formes quadratiques. Deuxième mémoire. Recherches sur les parallélloèdres
which drills tunnels through high mountains,” Technical Report No. 06-26, ZIB, primitifs,” J. Reine Angew. Math. 1908, 198–287.
Takustr., Berlin, 2006. 37
H. W. Kuhn, “The Hungarian method for the assignment problem,” Nav. Res.
21
H. Shen, J. Liu, and L. Fu, “Self-learning Monte Carlo with deep neural Logostic Q. 2(12), 83–97 (1955).
networks,” Phys. Rev. B 97, 205140 (2018). 38
M. R. Shirts and J. D. Chodera, “Statistically optimal analysis of samples from
22
R. Habib and D. Barber, “Auxiliary variational MCMC,” in International multiple equilibrium states,” J. Chem. Phys. 129(12), 124105 (2008).
Conference on Learning Representations (ICLR, La Jolla, CA, 2019). 39
H. A. Kramers, “Brownian motion in a field of force and the diffusion model of
23
J. Song, S. Zhao, and S. Ermon, “A-NICE-MC: Adversarial training for chemical reactions,” Physica 7(4), 284–304 (1940).
02 February 2024 01:29:53
MCMC,” in Advances in Neural Information Processing Systems, edited by 40
J. Hermann, Z. Schätzle, and F. Noé, “Deep neural network solution of the
I. Guyon, U. V. Luxburg, S. Bengio, H. Wallach, R. Fergus, S. Vish- electronic Schrödinger equation,” Nature Chem. 12, 891–897 (2020).
wanathan, and R. Garnett (Curran Associates, Inc., 2017), Vol. 30, pp. 41
K. Schütt, P.-J. Kindermans, H. E. Sauceda Felix, S. Chmiela, A. Tkatchenko,
5140–5150. and K.-R. Müller, “SchNet: A continuous-filter convolutional neural network for
24
D. Levy, M. D. Hoffman, and J. Sohl-Dickstein, “Generalizing Hamiltonian modeling quantum interactions,” in Advances in Neural Information Processing
Monte Carlo with neural networks,” in International Conference on Learning Systems (Curran Associates, Inc., 2017), pp. 991–1001.
Representations (ICLR, La Jolla, CA, 2018). 42
N. Thomas, T. Smidt, S. Kearnes, L. Yang, L. Li, K. Kohlhoff, and P. Riley, “Ten-
25
M. Dibak, L. Klein, and F. Noé, “Temperature-steerable flows,” arXiv:2012.00429 sor field networks: Rotation-and translation-equivariant neural networks for 3D
(2020). point clouds,” arXiv:1802.08219 (2018).
26 43
F. Noé, S. Olsson, J. Köhler, and H. Wu, “Boltzmann generators: Sampling J. Köhler, L. Klein, and F. Noé, “Equivariant flows: Exact likelihood generative
equilibrium states of many-body systems with deep learning,” Science 365(6457), learning for symmetric densities,” in Proceedings of the 37th International Con-
eaaw1147 (2019). ference on Machine Learning, edited by Hal Daumé III and Aarti Singh (PMLR,
27
L. Dinh, J. Sohl-Dickstein, and S. Bengio, “Density estimation using Real NVP,” 2020), Vol. 119, pp. 5361–5370.
arXiv:1605.08803 (2016). 44
D. P. Kingma and J. Ba, “Adam: A method for stochastic optimization,” in 3rd
28
D. Jimenez Rezende and S. Mohamed, “Variational inference with normalizing International Conference on Learning Representations, edited by Y. Bengio and
flows,” arXiv:1505.05770 (2015). Y. LeCun (ICLR, San Diego, CA, 2015).
J. Chem. Phys. 154, 074101 (2021); doi: 10.1063/5.0032346 154, 074101-8
© Author(s) 2021
You might also like
- College Physics 4th Edition Giambattista Test Bank 1Document31 pagesCollege Physics 4th Edition Giambattista Test Bank 1tracy100% (43)
- An Extension of Newton-Type Algorithms For Nonlinear Process ControlDocument6 pagesAn Extension of Newton-Type Algorithms For Nonlinear Process ControlGustavo Gabriel Jimenez100% (1)
- 220-12 Volt DC Power SupplyDocument16 pages220-12 Volt DC Power Supplychachunasayan89% (18)
- Temperature Dependence of Energy Transport in The Z Chiral Clock ModelDocument12 pagesTemperature Dependence of Energy Transport in The Z Chiral Clock ModelDaniele MarconiNo ratings yet
- Project ProposalDocument10 pagesProject ProposalPrathamNo ratings yet
- Andreas M. Lauchli - Numerical Simulations of Frustrated SystemsDocument34 pagesAndreas M. Lauchli - Numerical Simulations of Frustrated SystemsImaxSWNo ratings yet
- ChuangDocument8 pagesChuangKamal ChoudharyNo ratings yet
- Machine Learning Phases of Strongly Correlated FermionsDocument9 pagesMachine Learning Phases of Strongly Correlated FermionsalfonsoNo ratings yet
- King Et Al-2021-Nature CommunicationsDocument6 pagesKing Et Al-2021-Nature CommunicationsSteve KingNo ratings yet
- A Reduced Variational Approach For Searching Cycles in High-Dimensional SystemsDocument25 pagesA Reduced Variational Approach For Searching Cycles in High-Dimensional Systemsim.marzaNo ratings yet
- Meshless Methods1Document4 pagesMeshless Methods1herevenkatNo ratings yet
- Recognition of Temporal Sequences of Patterns Using State-Dependent SynapsesDocument18 pagesRecognition of Temporal Sequences of Patterns Using State-Dependent Synapsesyixinah516No ratings yet
- Entire Spectrum of Fully Many-Body Localized Systems Using Tensor NetworksDocument12 pagesEntire Spectrum of Fully Many-Body Localized Systems Using Tensor NetworksAnonymous lqvgUsVckNo ratings yet
- 1 s2.0 S0370269323004227 MainDocument7 pages1 s2.0 S0370269323004227 Mainsakoufah adrianNo ratings yet
- Reese JM Pure Coupled Continuum Hydrodynamics and Molecular Dynamics Method For Multiscale Simulation Aug 2011Document9 pagesReese JM Pure Coupled Continuum Hydrodynamics and Molecular Dynamics Method For Multiscale Simulation Aug 2011Avijit KarmakarNo ratings yet
- Clementi Diffussion MAps JCPDocument11 pagesClementi Diffussion MAps JCPDavid BeveridgeNo ratings yet
- Hard-Disk Equation of State: First-Order Liquid-Hexatic Transition in Two Dimensions With Three Simulation MethodsDocument8 pagesHard-Disk Equation of State: First-Order Liquid-Hexatic Transition in Two Dimensions With Three Simulation MethodsOzanBurakEriçokNo ratings yet
- Physical Quantum AlgorithmsDocument7 pagesPhysical Quantum AlgorithmsSankha DeepNo ratings yet
- Non-Markovian Quantum Dynamics in Strongly Coupled Multimode Cavities Conditioned On Continuous MeasurementDocument16 pagesNon-Markovian Quantum Dynamics in Strongly Coupled Multimode Cavities Conditioned On Continuous MeasurementMartta TervakoskiNo ratings yet
- 1104 5491v2 PDFDocument19 pages1104 5491v2 PDFJacky LaoNo ratings yet
- Quantum Monte CarloDocument6 pagesQuantum Monte Carlomar.sedanoortNo ratings yet
- Laurence E. Fried and Gregory S. Ezra - PERTURB: A Special-Purpose Algebraic Manipulation Program For Classical Perturbation TheoryDocument15 pagesLaurence E. Fried and Gregory S. Ezra - PERTURB: A Special-Purpose Algebraic Manipulation Program For Classical Perturbation TheoryOmasazzNo ratings yet
- Nino Garis - Numerical Solutions of The One Speed Neutron Transport Equation in Two Medium Slabs and Spheres (1991)Document52 pagesNino Garis - Numerical Solutions of The One Speed Neutron Transport Equation in Two Medium Slabs and Spheres (1991)Strawberry FieldsNo ratings yet
- RadevDocument3 pagesRadevIrenataNo ratings yet
- Umbrella Sampling - A ReviewDocument11 pagesUmbrella Sampling - A ReviewdanieloshkaNo ratings yet
- Decoherence ProblemDocument6 pagesDecoherence ProblemmrpcuNo ratings yet
- Spontaneous Symmetry Breaking in Replica Field Theory: 10.1103/physrevd.96.065012Document14 pagesSpontaneous Symmetry Breaking in Replica Field Theory: 10.1103/physrevd.96.065012Angie DamarisNo ratings yet
- Andrew K. Henrick, Tariq D. Aslam and Joseph M. Powers - Simulations of Pulsating One-Dimensional Detonations With True Fifth Order AccuracyDocument19 pagesAndrew K. Henrick, Tariq D. Aslam and Joseph M. Powers - Simulations of Pulsating One-Dimensional Detonations With True Fifth Order AccuracyNikeShoxxxNo ratings yet
- 1609 01552 PDFDocument12 pages1609 01552 PDFGNo ratings yet
- 0907 2948 PDFDocument17 pages0907 2948 PDFCarlos LopezNo ratings yet
- Certifying Multilevel Coherence in The Motional State of A Trapped IonDocument14 pagesCertifying Multilevel Coherence in The Motional State of A Trapped IonNikhilKotibhaskarNo ratings yet
- Data-Driven Computation of Molecular Reaction Coordinates: Andreas Bittracher, Ralf Banisch, and Christof SCH UtteDocument24 pagesData-Driven Computation of Molecular Reaction Coordinates: Andreas Bittracher, Ralf Banisch, and Christof SCH Uttehuevonomar05No ratings yet
- Boost-Invariant (2+1) - Dimensional Anisotropic Hydrodynamics: PACS Numbers: 12.38.Mh, 24.10.Nz, 25.75.Ld, 25.75.-qDocument44 pagesBoost-Invariant (2+1) - Dimensional Anisotropic Hydrodynamics: PACS Numbers: 12.38.Mh, 24.10.Nz, 25.75.Ld, 25.75.-qsharky84No ratings yet
- Decoherence Limits The Cost To Simulate An Anharmonic OscillatorDocument21 pagesDecoherence Limits The Cost To Simulate An Anharmonic OscillatorSRINIDHI PAWARNo ratings yet
- Numerical Atomic Orbitals For Linear-Scaling Calculations: 共Received 6 April 2001; Published 28 November 2001兲Document9 pagesNumerical Atomic Orbitals For Linear-Scaling Calculations: 共Received 6 April 2001; Published 28 November 2001兲AytaçÇelikNo ratings yet
- Distributed Quantum Phase Estimation With Entangled PhotonsDocument8 pagesDistributed Quantum Phase Estimation With Entangled Photons박정우No ratings yet
- Phase Equilibria in The Polydisperse Zwanzig Model of Hard RodsDocument14 pagesPhase Equilibria in The Polydisperse Zwanzig Model of Hard RodsGilberto RuizNo ratings yet
- Sequence Space JacobianDocument84 pagesSequence Space JacobianStan ChrisinskyNo ratings yet
- QMMM MethodDocument8 pagesQMMM MethodSubarnaNo ratings yet
- Load Balancing Strategies For The DSMC Simulation of Hypersonic Flows Using HPCDocument12 pagesLoad Balancing Strategies For The DSMC Simulation of Hypersonic Flows Using HPCnanilaurntiuNo ratings yet
- Modelos DFTDocument17 pagesModelos DFTAlberto Núñez CardezoNo ratings yet
- Non Linear CH PaperDocument9 pagesNon Linear CH PaperseenuNo ratings yet
- Statement of Research Interests-Example 3Document7 pagesStatement of Research Interests-Example 3NedelcuGeorgeNo ratings yet
- Collective Variable-Based Enhanced Sampling and Machine Learning PDFDocument17 pagesCollective Variable-Based Enhanced Sampling and Machine Learning PDFwenhuishenNo ratings yet
- Nonlinear Heat Transport in Mesoscopic Conductors: Rectification, Peltier Effect and Wiedemann-Franz LawDocument8 pagesNonlinear Heat Transport in Mesoscopic Conductors: Rectification, Peltier Effect and Wiedemann-Franz LawJeanette AmayaNo ratings yet
- An Adaptive Variational Algorithm For Exact Molecular Simulations On A Quantum ComputerDocument9 pagesAn Adaptive Variational Algorithm For Exact Molecular Simulations On A Quantum ComputerRaiyan RahmanNo ratings yet
- 07 - chapter 2 مهم جدا جدا لحساب activation energyDocument30 pages07 - chapter 2 مهم جدا جدا لحساب activation energymahmoudNo ratings yet
- Micadei - Experimental Rectification of Entropy Production by A Maxwell's Demon in A Quantum SystemDocument10 pagesMicadei - Experimental Rectification of Entropy Production by A Maxwell's Demon in A Quantum SystemFábio LucasNo ratings yet
- Numerical Algorithms For The Hypercube Concu - 1988 - Mathematical and ComputerDocument3 pagesNumerical Algorithms For The Hypercube Concu - 1988 - Mathematical and ComputerdarshanNo ratings yet
- The Emergent Copenhagen Interpretation of Quantum Mechanics - Timothy J. HollowoodDocument21 pagesThe Emergent Copenhagen Interpretation of Quantum Mechanics - Timothy J. HollowoodCambiador de MundoNo ratings yet
- Coherent Feedback Cooling of A Nanomechanical Membrane With Atomic SpinsDocument13 pagesCoherent Feedback Cooling of A Nanomechanical Membrane With Atomic Spinswork outNo ratings yet
- GPU-based Reaction Ensemble Monte Carlo Method For Non-Ideal Plasma ThermodynamicsDocument28 pagesGPU-based Reaction Ensemble Monte Carlo Method For Non-Ideal Plasma ThermodynamicsAndreina Dell'olioNo ratings yet
- Mesoscopic Quantum Coherences in Cavity QED: Preparation and Decoherence Monitoring SchemesDocument15 pagesMesoscopic Quantum Coherences in Cavity QED: Preparation and Decoherence Monitoring SchemesVictor Viera CastilloNo ratings yet
- 123Document4 pages123vato otavNo ratings yet
- On The Mathematical Foundations of Diffusion Monte Carlo: Michel Caffarel, Pierre Del Moral and Luc de MontellaDocument38 pagesOn The Mathematical Foundations of Diffusion Monte Carlo: Michel Caffarel, Pierre Del Moral and Luc de MontellaPinakiNo ratings yet
- E (3) - Equivariant Graph Neural Networks For Data-Ef Ficient and Accurate Interatomic PotentialsDocument11 pagesE (3) - Equivariant Graph Neural Networks For Data-Ef Ficient and Accurate Interatomic PotentialsichbinkeinrobotjaNo ratings yet
- Paper On Umbrella SamplingDocument11 pagesPaper On Umbrella SamplingSayed Abu SufyanNo ratings yet
- Mathematics 09 01442Document12 pagesMathematics 09 01442Golam ZakariaNo ratings yet
- (1959) Studies in Molecular Dynamics. I. General MethodDocument8 pages(1959) Studies in Molecular Dynamics. I. General MethodJakler NicheleNo ratings yet
- Polarization Coding of Quantum Information (Optics Communications, Vol. 123, Issue 1-3) (1996)Document10 pagesPolarization Coding of Quantum Information (Optics Communications, Vol. 123, Issue 1-3) (1996)sepot24093No ratings yet
- Rodrguez Her N Ndez 2014Document6 pagesRodrguez Her N Ndez 2014Tengku Arief Buana PerkasaNo ratings yet
- Journal of Quantitative Spectroscopy & Radiative Transfer: Laurent Soucasse, Philippe Riviere, Anouar SoufianiDocument9 pagesJournal of Quantitative Spectroscopy & Radiative Transfer: Laurent Soucasse, Philippe Riviere, Anouar Soufianimbb appNo ratings yet
- (W-7443) Differentiable Molecular Simulations For Control and LearningDocument16 pages(W-7443) Differentiable Molecular Simulations For Control and LearningNazmul SaqibNo ratings yet
- (W-6135) (2023) Accelerated Sampling of Rare Events Using A Neural Network Bias PotentialDocument12 pages(W-6135) (2023) Accelerated Sampling of Rare Events Using A Neural Network Bias PotentialNazmul SaqibNo ratings yet
- (W-6725) TensorMol - NN Augmented With Long Ramge PhysicsDocument9 pages(W-6725) TensorMol - NN Augmented With Long Ramge PhysicsNazmul SaqibNo ratings yet
- (W-6597) Deep-Neural-Network Approach To Solving The Ab Initio Nuclear Structure ProblemDocument9 pages(W-6597) Deep-Neural-Network Approach To Solving The Ab Initio Nuclear Structure ProblemNazmul SaqibNo ratings yet
- How To Do Machine Learning With Small DataDocument22 pagesHow To Do Machine Learning With Small DataNazmul SaqibNo ratings yet
- Series EA Electric Actuators Low Torque, Medium Torque, and Spring Return Manual 1321-In-003!0!13Document16 pagesSeries EA Electric Actuators Low Torque, Medium Torque, and Spring Return Manual 1321-In-003!0!13Isaac MonterreyNo ratings yet
- Arches Vaults and Dome FinalDocument39 pagesArches Vaults and Dome FinalPratima Mahesh0% (1)
- 096 09247 03 - TDSDocument2 pages096 09247 03 - TDSMuhamad Samsul BahriNo ratings yet
- WCM - Yamashina - 05 - Professional Maintenance - Chapter - 3 - ADocument100 pagesWCM - Yamashina - 05 - Professional Maintenance - Chapter - 3 - ACíceroNo ratings yet
- Rice 1788Document70 pagesRice 1788Dr. Rohit SinghNo ratings yet
- Jurnal Fatimah SimbolonDocument8 pagesJurnal Fatimah SimbolonFatimah SimbolonNo ratings yet
- tp2 Systc3a8mes Asservis Numc3a9riquesDocument1 pagetp2 Systc3a8mes Asservis Numc3a9riquesomarNo ratings yet
- CHT307 - Ktu QbankDocument3 pagesCHT307 - Ktu QbankYuxin CasioNo ratings yet
- CBSE Class 12 Chemistry Quick Revision Notes Co-Ordination CompoundsDocument8 pagesCBSE Class 12 Chemistry Quick Revision Notes Co-Ordination CompoundsAbid waniNo ratings yet
- Ip Rating PDFDocument1 pageIp Rating PDFGurunadha Rao RamachandraNo ratings yet
- Isolated Footing DesignDocument106 pagesIsolated Footing DesignAlemayehu Miteku100% (1)
- Microscope Laboratory Activities 2023Document7 pagesMicroscope Laboratory Activities 2023Izzy Dale Cagas SilangaNo ratings yet
- Module 4 Work Energy PowerDocument7 pagesModule 4 Work Energy PowerGreen BrainNo ratings yet
- Gujarat Technological UniversityDocument1 pageGujarat Technological UniversityBhagirath sinh ZalaNo ratings yet
- Fujifilm Eluxeo BL 7000Document78 pagesFujifilm Eluxeo BL 7000AngeloNo ratings yet
- RadioaktivlikDocument3 pagesRadioaktivlikgultaczaurNo ratings yet
- Synthesis of Zns / Sodium Hexameta Phosphate Nanoparticles: R. Mohan, B. Rajamannan and S. SankarrajanDocument6 pagesSynthesis of Zns / Sodium Hexameta Phosphate Nanoparticles: R. Mohan, B. Rajamannan and S. SankarrajanphysicsjournalNo ratings yet
- Current Logic 1000V - 24V Power SupplyDocument2 pagesCurrent Logic 1000V - 24V Power Supplyraza239No ratings yet
- AC Induction Motor MC WebinarDocument61 pagesAC Induction Motor MC WebinarAdeNo ratings yet
- Cape Wind 462 MW GenerationDocument78 pagesCape Wind 462 MW GenerationmilanovsdNo ratings yet
- EE6512-Electrical Machines LaboratoryDocument82 pagesEE6512-Electrical Machines LaboratoryGopinath B L NaiduNo ratings yet
- s4 Physics Paper 3 Exam 5Document4 pagess4 Physics Paper 3 Exam 5MUSOKE GERALDNo ratings yet
- Math Reviewer Lesson 1 3Document27 pagesMath Reviewer Lesson 1 3Junel BitangaNo ratings yet
- Pamantasan NG Lungsod NG ValenzuelaDocument2 pagesPamantasan NG Lungsod NG ValenzuelaDiane de OcampoNo ratings yet
- Mechanics of Fluids Si Edition 5th Edition Potter Test BankDocument3 pagesMechanics of Fluids Si Edition 5th Edition Potter Test Bankwalleyeemploye2g100% (28)
- ENE - Combined Updated For Use of AY2020Document2 pagesENE - Combined Updated For Use of AY2020Alzheimer RachNo ratings yet
- 425ºC AHT: Ower EATDocument2 pages425ºC AHT: Ower EATSivagurunathan SpNo ratings yet